A Fast Sampling Method of Exploring Graphlet Degrees of Large Directed and Undirected Graphs
Exploring small connected and induced subgraph patterns (CIS patterns, or graphlets) has recently attracted considerable attention. Despite recent efforts on computing the number of instances a specific graphlet appears in a large graph (i.e., the total number of CISes isomorphic to the graphlet), little attention has been paid to characterizing a node’s graphlet degree, i.e., the number of CISes isomorphic to the graphlet that include the node, which is an important metric for analyzing complex networks such as social and biological networks. Similar to global graphlet counting, it is challenging to compute node graphlet degrees for a large graph due to the combinatorial nature of the problem. Unfortunately, previous methods of computing global graphlet counts are not suited to solve this problem. In this paper we propose sampling methods to estimate node graphlet degrees for undirected and directed graphs, and analyze the error of our estimates. To the best of our knowledge, we are the first to study this problem and give a fast scalable solution. We conduct experiments on a variety of real-word datasets that demonstrate that our methods accurately and efficiently estimate node graphlet degrees for graphs with millions of edges.
💡 Research Summary
The paper tackles the problem of estimating node‑level graphlet (also called motif or connected induced subgraph, CIS) degrees in large graphs, a task that has received far less attention than global graphlet counting. A node’s graphlet degree vector records, for each automorphism orbit of a graphlet, how many instances of that graphlet contain the node in that specific position. These vectors are powerful descriptors for applications ranging from protein function prediction to link prediction in social networks, but exact enumeration is infeasible for massive graphs because the number of 3‑ and 4‑node CISes that involve a high‑degree node can easily exceed 10¹⁴.
To overcome this combinatorial explosion, the authors propose a two‑stage sampling‑based framework.
Stage 1 – Targeted sampling of 3‑ and 4‑node CISes.
Six specialized samplers are introduced: Randgraf‑3‑1, Randgraf‑3‑2, Randgraf‑4‑1, Randgraf‑4‑2, Randgraf‑4‑3, and Randgraf‑4‑4. Each sampler generates a CIS that must contain a given query node v, but each is limited to a subset of the possible orbits.
- Randgraf‑3‑1* picks two distinct neighbors of v uniformly at random, forming a triangle (or a 3‑node star). It only contributes to undirected orbits 2 and 3, and its sampling probability is p(3,1)₂ = p(3,1)₃ = 1/φᵥ where φᵥ = dᵥ(dᵥ − 1)/2.
- Randgraf‑3‑2* selects a neighbor u of v with probability proportional to (dᵤ − 1) (denoted α(v)ᵤ), then picks a random neighbor w of u distinct from v. This sampler contributes to orbits 1 and 3 with probabilities p(3,2)₁ = 1/ϕᵥ and p(3,2)₃ = 2/ϕᵥ, where ϕᵥ = ∑_{u∈N(v)}(dᵤ − 1).
The four 4‑node samplers extend the same idea: they combine uniform and weighted selections among v’s neighbors, v’s neighbor‑of‑neighbor sets, and sometimes allow the third sampled node to coincide with the second, thereby also covering 3‑node CISes. Each method’s bias (the probability of landing in a particular orbit) is derived analytically (e.g., Randgraf‑4‑1 has Φ¹ᵥ = (dᵥ − 1)·ϕᵥ). Because the bias is known exactly, an unbiased estimator can be obtained simply by dividing the observed count by the corresponding probability.
Stage 2 – Unbiased estimation of orbit degrees.
For undirected graphs the authors build the SAND (Sampling for ANalysis of graphlet Degrees) framework. For each orbit i (i = 0…14) they run a collection of independent samplers, each producing an unbiased estimate ˆcᵢⱼ of the true orbit count dᵢ(v). Theorem 2 (combining unbiased estimators) tells us that the minimum‑variance linear combination is ˆdᵢ(v) = ∑ⱼ αⱼ·ˆcᵢⱼ where αⱼ ∝ 1/Var(ˆcᵢⱼ). The resulting variance is 1/∑ⱼ 1/Var(ˆcᵢⱼ), which can be computed directly from the known sampling probabilities and the number of draws Kⱼ allocated to each sampler. This gives a closed‑form expression for the confidence interval of each orbit degree, allowing practitioners to decide a priori how many samples are needed to achieve a desired error ε with confidence 1 − δ (via a standard Chernoff‑type bound).
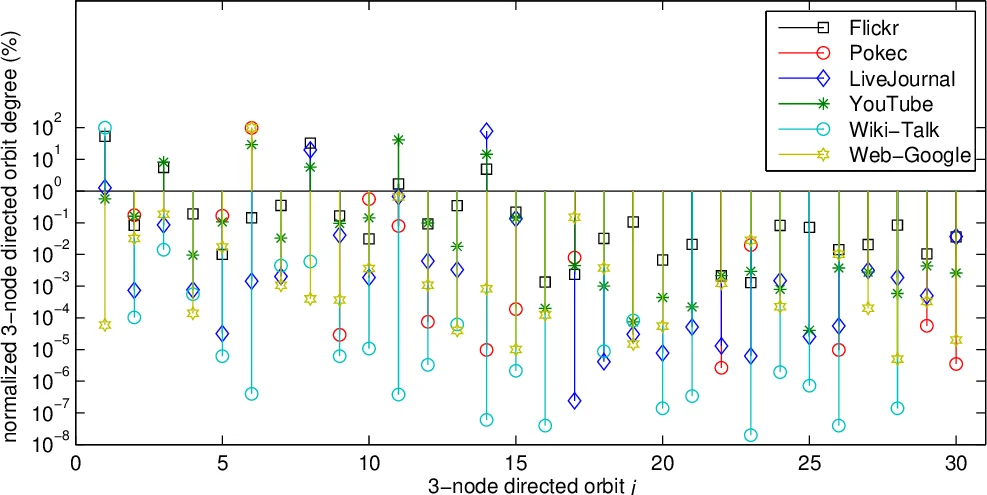
For directed graphs the authors extend the approach to SAND‑3D, which handles the 13 three‑node directed graphlets and their 30 distinct orbits. The same sampling primitives are used, but the direction labels (→, ←, ↔) are preserved, and the bias formulas are adapted accordingly.
Theoretical contributions.
The paper formalizes the sampling problem as estimating the cardinalities of non‑overlapping subsets S₁,…,Sᵣ of a larger set S (the set of all CISes containing v). Theorem 1 provides an unbiased estimator for each |Sᵢ| given a sampling distribution with known probabilities pᵢ, together with variance and covariance formulas. By plugging the analytically derived pᵢ for each sampler, the authors obtain exact variance expressions for every orbit estimator. This rigorous statistical foundation distinguishes the work from heuristic sampling methods that lack error guarantees.
Experimental evaluation.
The authors evaluate SAND and SAND‑3D on twelve publicly available networks, including social graphs (e.g., Wiki‑Talk, YouTube), citation graphs (DBLP), and biological interaction networks. They compare against state‑of‑the‑art exact enumerators such as Orca and PGD. Key findings:
- Accuracy – Mean absolute error (MAE) across all orbits is typically below 0.01, indicating that the sampled estimates are virtually indistinguishable from exact counts.
- Speed – The sampling‑based methods are 10× to 200× faster than exhaustive enumeration, with the speedup growing for nodes of higher degree because the sampling cost scales linearly with degree while enumeration scales combinatorially.
- Scalability – The methods comfortably handle graphs with millions of edges and nodes; memory overhead remains O(1) beyond the storage of the input graph.
- Utility – Using the estimated orbit degree vectors for downstream tasks (node clustering, similarity search, anomaly detection) yields performance comparable to or better than using exact vectors, confirming that the small estimation error does not degrade practical usefulness.
Impact and future directions.
By delivering a fast, provably accurate, and memory‑efficient solution for node‑level graphlet degree estimation, the paper fills a critical gap in network science toolkits. The approach enables large‑scale analyses that were previously limited to global motif counts, opening the door to richer node embeddings, more precise role‑based clustering, and real‑time monitoring of evolving networks. Potential extensions include (i) handling larger graphlets (5‑node or more) via hierarchical sampling, (ii) integrating the estimators into graph neural network pipelines as additional structural features, and (iii) adapting the framework to dynamic graphs where orbit degrees must be updated incrementally as edges arrive or disappear.
In summary, the work presents a well‑theorized, practically validated, and broadly applicable methodology for estimating node graphlet orbit degrees in both undirected and directed massive graphs, representing a significant advancement for graph mining and network analysis.
Comments & Academic Discussion
Loading comments...
Leave a Comment