A Bio-Synthetic Modulator Model for Diffusion-based Molecular Communications
In diffusion-based molecular communication (DMC), one important functionality of a transmitter nano-machine is signal modulation. In particular, the transmitter has to be able to control the release of signaling molecules for modulation of the information bits. An important class of control mechanisms in natural cells for releasing molecules is based on ion channels which are pore-forming proteins across the cell membrane whose opening and closing may be controlled by a gating parameter. In this paper, a modulator for DMC based on ion channels is proposed which controls the rate at which molecules are released from the transmitter by modulating a gating parameter signal. Exploiting the capabilities of the proposed modulator, an on-off keying modulation scheme is introduced and the corresponding average modulated signal, i.e., the average release rate of the molecules from the transmitter, is derived in the Laplace domain. By making a simplifying assumption, a closed-form expression for the average modulated signal in the time domain is obtained which constitutes an upper bound on the total number of released molecules regardless of this assumption. The derived average modulated signal is compared to results obtained with a particle based simulator. The numerical results show that the derived upper bound is tight if the number of ion channels distributed across the transmitter (cell) membrane is small.
💡 Research Summary
This paper proposes a novel bio-synthetic modulator model for Diffusion-based Molecular Communication (DMC) systems, drawing inspiration from the molecular release mechanisms of natural cells. The core idea is to utilize ion channels—pore-forming membrane proteins whose opening and closing is controlled by a gating parameter (e.g., voltage)—as the foundation for a synthetic transmitter.
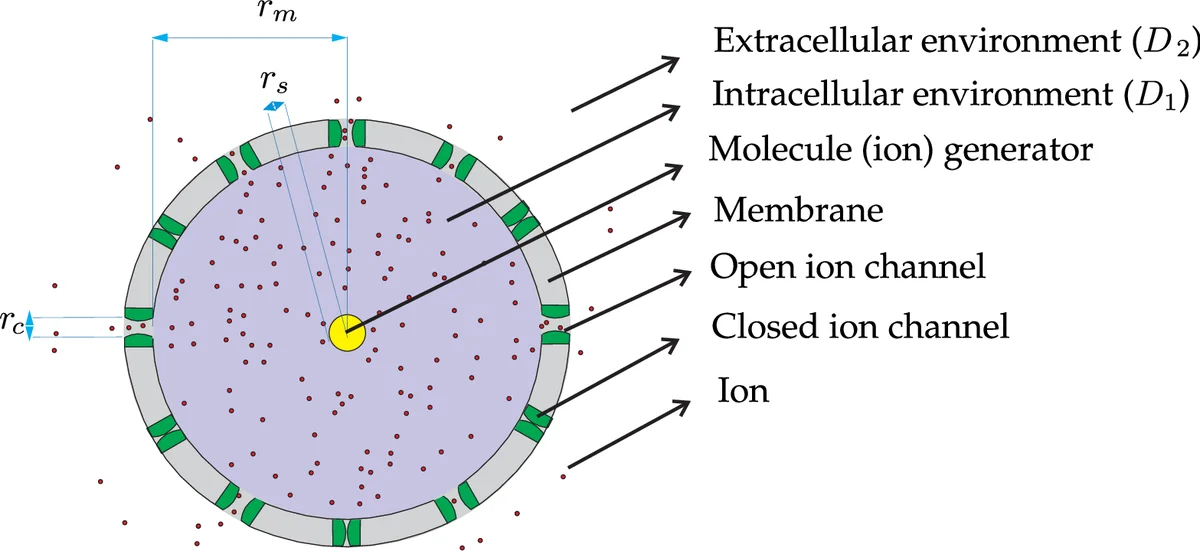
The authors model the transmitter as a spherical cell of radius r_m, whose membrane is uniformly embedded with N voltage-gated ion channels. The signaling molecules (ions) are produced at a constant rate by a central organelle inside the cell until an internal concentration threshold T is reached. The diffusion constants inside and outside the cell are D1 and D2, respectively. A key modeling step is the introduction of the “average membrane permeability” z, defined as the ratio of the total open channel area to the total membrane area (z = N π r_c^2 / (4 π r_m^2)). This allows the treatment of the stochastic channel openings as a uniformly permeable boundary.
Exploiting this modulator, the paper introduces a simple time-slotted On-Off Keying (OOK) modulation scheme. To transmit bit ‘0’, a voltage V_off is applied to keep the ion channels mostly closed throughout the symbol interval T. To transmit bit ‘1’, a voltage V_on is applied for the first T1 seconds to open the channels, followed by V_off for the remaining T2 = T - T1 seconds to close them and allow for internal molecule replenishment and channel clearing to mitigate Inter-Symbol Interference (ISI).
The primary analytical contribution is the derivation of the average modulated signal, i.e., the average molecule release rate from the transmitter, for the proposed IBM under the OOK scheme for bit ‘1’. This is achieved by solving the corresponding diffusion problem with boundary conditions that incorporate the average permeability z. The solution yields a closed-form expression for the Laplace transform of the average release rate, Ḡ(s). In general, obtaining the time-domain signal requires numerical inverse Laplace transform.
By making a simplifying assumption of zero concentration outside the cell membrane (C2=0), the authors derive a closed-form expression for the average release rate in the time domain, denoted as g_UB(t). They rigorously prove that the time integral of g_UB(t) provides an upper bound on the total number of molecules released, regardless of the validity of the C2=0 assumption.
The theoretical analysis is validated using a Particle-Based Simulator (PBS). The numerical results demonstrate two key findings: 1) The derived upper bound g_UB(t) is very tight to the actual simulated release rate when the number of ion channels N (and thus the permeability z) is small. This is because a low release rate maintains a low external concentration, making the C2=0 assumption realistic. 2) The shape of the release signal is predominantly determined by the diffusion process of molecules inside the cell, while the electrical dynamics of the ion channels (characterized by their opening/closing time constant t_c) have a negligible effect for typical parameters. This implies that the fundamental bandwidth limitation of such a modulator stems from the diffusion process itself, not the channel gating electronics.
In conclusion, this work bridges biological mechanisms and engineering design for DMC by providing a realistic, analytically tractable modulator model. It offers valuable insights into the physical constraints of synthetic molecular transmitters and provides a useful upper-bound tool for system design. The model establishes a foundation for future work exploring more complex modulation schemes, different gating mechanisms (e.g., ligand-gated channels), and integration with receiver models.
Comments & Academic Discussion
Loading comments...
Leave a Comment