Intriguing symmetry in statistical structures of Siberian larch transcriptome
The paper presents a novel approach to infer a structuredness in a set of symbol sequences such as transcriptome nucleotide sequences. A distribution pattern of triplet frequencies in the Siberian larch (\textit{Larix sibirica}~Ledeb.) transcriptome sequences was investigated in the presented study. It was found that the larch transcriptome demonstrates a number of unexpected symmetries in the statistical and combinatorial properties.
💡 Research Summary
The study investigates statistical regularities and unexpected symmetries in the transcriptome of Siberian larch (Larix sibirica) by converting nucleotide sequences into frequency dictionaries of triplets (3‑mers). From an initial set of 43,686 transcript sequences, only those longer than 200 bp were retained, resulting in 1,436 sequences for detailed analysis. Each sequence was transformed into a 64‑dimensional vector representing the normalized frequencies of all possible triplets (4³ = 64). These vectors define points in a Euclidean metric space, allowing quantitative comparison of sequences via Euclidean distance.
Clustering was performed using the K‑means algorithm with the number of clusters (K) varied from 2 to 5. For each K, 350 independent runs were executed to assess stability; a clustering was deemed stable when at least 95 % of runs produced the same partition. The most robust solution emerged at K = 3, where the inter‑cluster distances exceeded the sum of the respective cluster radii, confirming clear separation.
A key observation concerns Chargaff’s second parity rule, which predicts near‑equality of complementary base pair frequencies (e.g., A≈T, C≈G) within a single strand. When the two main clusters were examined separately, each displayed a relatively high deviation from this rule. However, when the clusters were combined, the deviation collapsed to near zero. This suggests that the transcriptome contains complementary sequences from both the (+) and (‑) strands in roughly equal proportions. To validate this hypothesis, BLAST searches were conducted to assign strand orientation to each transcript, confirming a balanced representation of both strands.
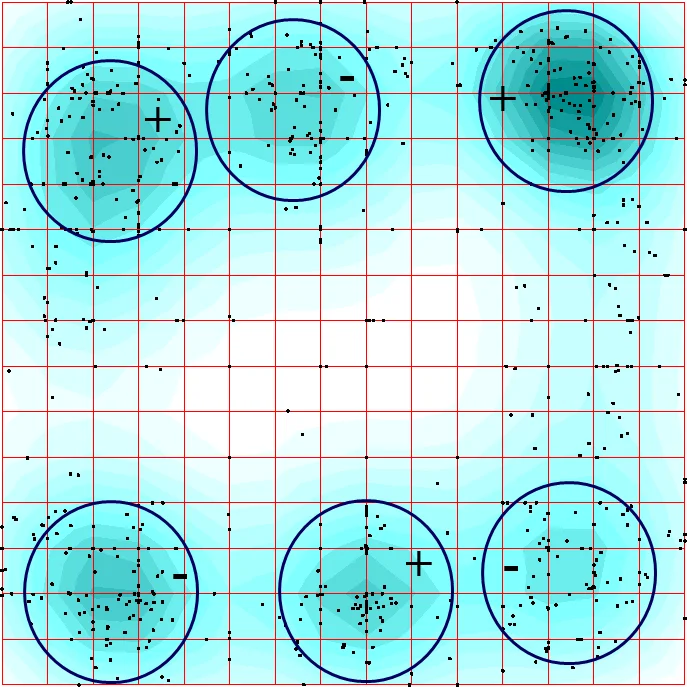
In addition to K‑means, the authors applied the Elastic Map technique, a manifold‑learning method that projects high‑dimensional data onto a two‑dimensional elastic surface. After initializing the surface with the first two principal components, points were connected to the surface by elastic springs and the surface was iteratively deformed to minimize total potential energy. Local density was visualized using Gaussian kernels, producing a colored map that revealed a striking octahedral arrangement: six distinct vertices (or “corners”) connected by edges, forming an octahedron in the projected space. This pattern differs from previously reported seven‑cluster structures in other transcriptomes, indicating a novel symmetry specific to the Siberian larch data.
The authors conclude that triplet‑frequency dictionaries provide a powerful framework for uncovering hidden structure in nucleotide sequence collections. The near‑cancellation of Chargaff‑rule violations across clusters supports the notion of symmetric expression from both DNA strands, while the octahedral geometry uncovered by Elastic Map visualisation points to a higher‑order combinatorial symmetry in the transcriptome. These findings not only introduce a new analytical pipeline for transcriptomic data but also offer statistical evidence for balanced bidirectional transcription in Siberian larch, with potential implications for understanding genome organization and expression regulation in conifers.
Comments & Academic Discussion
Loading comments...
Leave a Comment