Estimating the number of tissue resident macrophages
I provide a simple estimation for the number of macrophages in a tissue, arising from the hypothesis that they should keep infections below a certain threshold, above which neutrophils are recruited from blood circulation. The estimation reads Nm=a Ncel^{\alpha}/Nmax, where a is a numerical coefficient, the exponent {\alpha} is near 2/3, and Nmax is the maximal number of pathogens a macrophage may engulf in the time interval, tr, between pathogen replications.
💡 Research Summary
In this paper the author proposes a minimalist quantitative framework for estimating the number of resident macrophages (Nm) in a given tissue based on the idea that macrophages must keep the local pathogen load below a threshold that would otherwise trigger recruitment of circulating neutrophils. The model starts with the classic exponential growth law for bacteria in the early phase of infection, Np = Np(0) exp(t ln 2/ tr), where tr is the bacterial replication time (≈1 h). As the bacterial colony expands, the author argues that geometric constraints cause only the cells at the surface of the cluster to divide, because interior cells become space‑limited. Assuming a roughly spherical cluster, the radius scales as R ∼ Np^{1/3} and the surface area as R² ∼ Np^{2/3}. Consequently, the number of new bacteria generated during one replication interval is proportional to the surface area: ΔNp ≈ a0 Np^{2/3}, where a0 is a proportionality constant that captures details of the bacterial species and the tissue micro‑environment.
Macrophages are assumed to be uniformly distributed throughout the tissue and to possess high motility, so that within a few replication cycles (3–4 tr) the majority of them can be mobilized to the infection site. The immune system is considered successful if the number of newly produced bacteria (ΔNp) is smaller than the total number that can be eliminated by the macrophages present at the site. If N0m denotes the number of macrophages that have already arrived, and Nmax the maximal number of bacteria a single macrophage can engulf in one replication interval, the condition reads a0 Np^{2/3} < N0m · Nmax. Since N0m cannot exceed the total resident macrophage pool Nm, the inequality simplifies to a0 Np^{2/3} < Nm · Nmax.
To connect the pathogen count to a measurable tissue property, the author replaces Np by the total number of tissue cells Ncel, arguing that infection becomes clinically relevant when the fraction of injured cells exceeds a small, fixed proportion (e.g., 1 %). This substitution yields the central scaling law
Nm > a Ncel^{α} / Nmax,
where α≈2/3 reflects the surface‑to‑volume relationship of a three‑dimensional cluster, and a is a new dimensionless coefficient that incorporates the injury‑fraction threshold and the original geometric constant a0. The author treats Nmax as roughly constant across tissues (≈30–50 bacteria per hour for a hyper‑activated macrophage) and focuses on variations in a and Ncel.
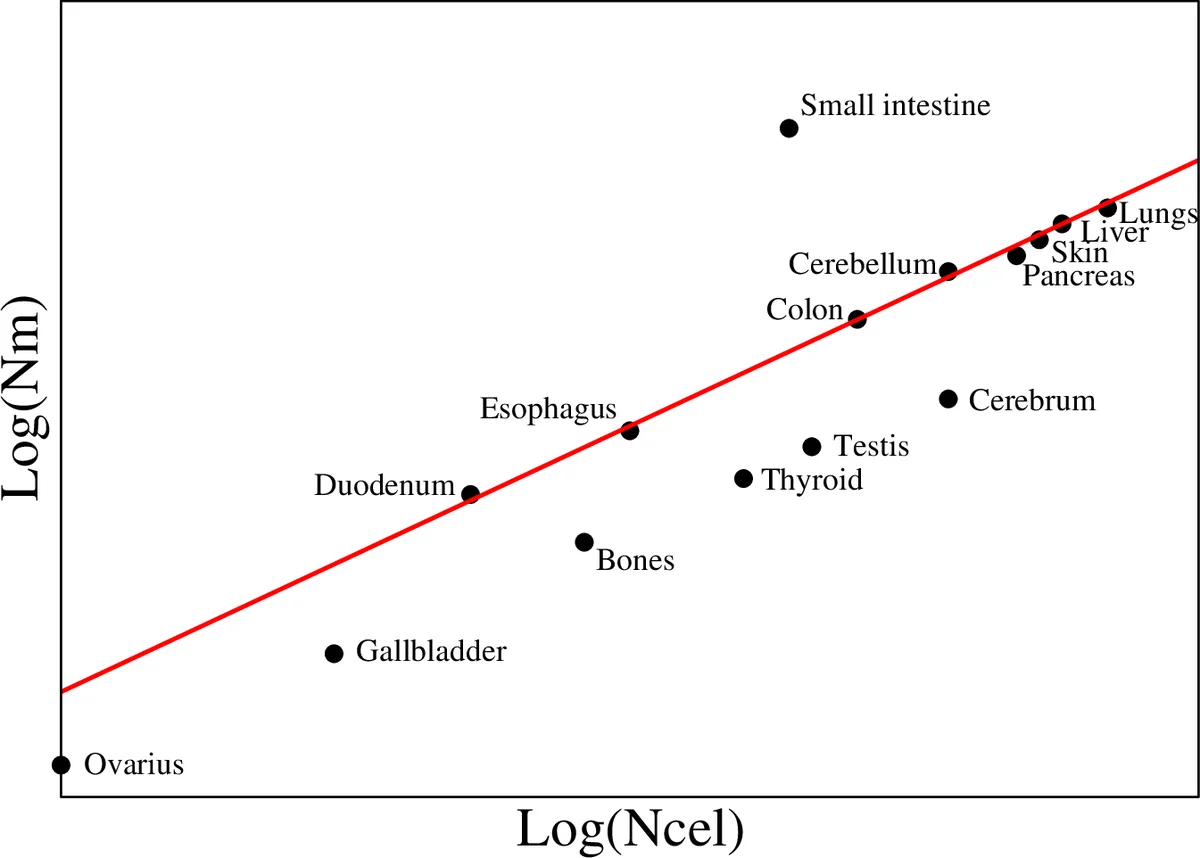
Taking logarithms gives log Nm > α log Ncel + log(a/Nmax). In a log‑log plot of Nm versus Ncel, “normal” tissues should line up along a straight line of slope α≈2/3. The author illustrates this with data from a previous study (Ref
Comments & Academic Discussion
Loading comments...
Leave a Comment