Scalable Ontological Query Processing over Semantically Integrated Life Science Datasets using MapReduce
To address the requirement of enabling a comprehensive perspective of life-sciences data, Semantic Web technologies have been adopted for standardized representations of data and linkages between data. This has resulted in data warehouses such as UniProt, Bio2RDF, and Chem2Bio2RDF, that integrate different kinds of biological and chemical data using ontologies. Unfortunately, the ability to process queries over ontologically-integrated collections remains a challenge, particularly when data is large. The reason is that besides the traditional challenges of processing graph-structured data, complete query answering requires inferencing to explicate implicitly represented facts. Since traditional inferencing techniques like forward chaining are difficult to scale up, and need to be repeated each time data is updated, recent focus has been on inferencing that can be supported using database technologies via query rewriting. However, due to the richness of most biomedical ontologies relative to other domain ontologies, the queries resulting from the query rewriting technique are often more complex than existing query optimization techniques can cope with. This is particularly so when using the emerging class of cloud data processing platforms for big data processing due to some additional overhead which they introduce. In this paper, we present an approach for dealing such complex queries on big data using MapReduce, along with an evaluation on existing real-world datasets and benchmark queries.
💡 Research Summary
The paper addresses the challenge of executing SPARQL queries over large, ontology‑rich life‑science RDF datasets such as UniProt, Bio2RDF, and Chem2Bio2RDF. Traditional forward‑chaining inference is infeasible at the terabyte scale because it requires materializing all inferred triples and must be repeated after each data update. Consequently, recent research has focused on query rewriting: expanding the original query using ontology axioms into a Union of Conjunctive Queries (UCQ). While this approach avoids data‑side materialization, biomedical ontologies are exceptionally deep and broad (e.g., UniProt’s class hierarchy exceeds 25 levels and contains over 1.3 million classes). As a result, rewritten UCQs often contain hundreds of UNION branches and thousands of JOINs, far exceeding the capabilities of conventional relational optimizers.
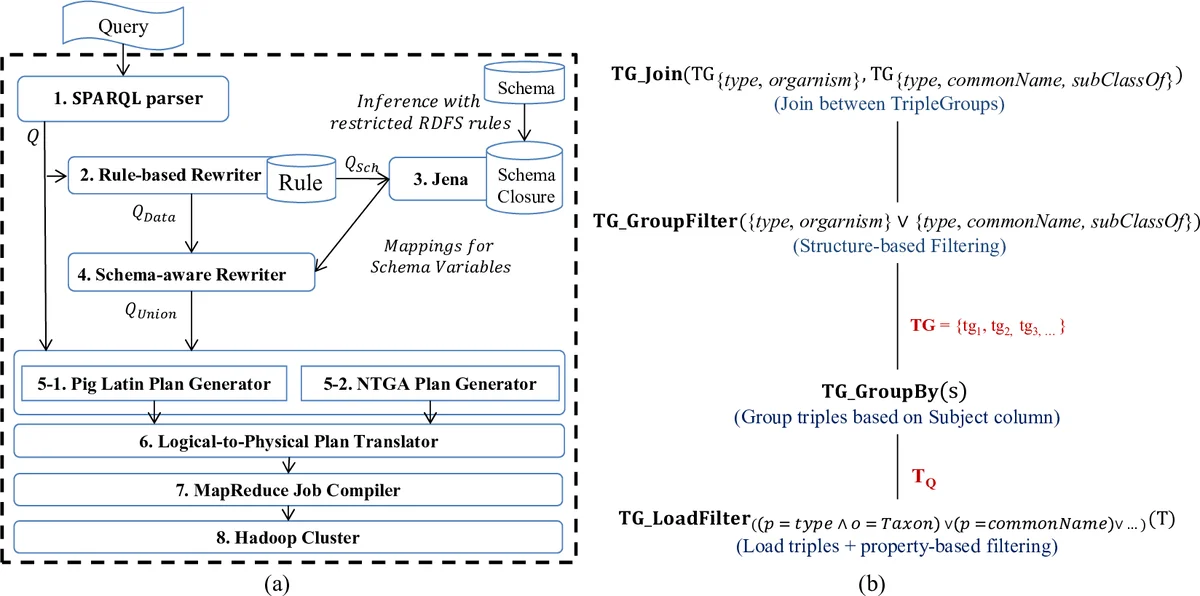
The authors propose a novel data model and algebra, the Nested Triple Group Data Model and Algebra (NTGA), to mitigate these problems in a MapReduce environment. NTGA treats sets of triples that share the same subject as first‑class “triple groups.” Three core operators are introduced: TG GroupBy, which groups all triples by subject in a single MapReduce job; TG GroupFilter, which discards groups that do not satisfy any star‑pattern of the query; and TG Join, which combines matching groups across different star‑patterns. By processing all star‑patterns simultaneously, the number of required MapReduce cycles drops from the naïve 2 n − 1 (where n is the number of star patterns) to n + 1 (one initial grouping/filtering job plus at most one join per star pattern). This reduction dramatically cuts the amount of data shuffled between map and reduce phases and lowers overall job latency.
To operationalize NTGA, the authors extend Apache Pig with a system called RAPID+. The workflow is as follows: (1) a SPARQL query is parsed via Jena ARQ or an extended Pig Latin interface; (2) a rule‑based query rewriter expands the query according to RDFS/OWL rules, producing a UCQ; (3) the UCQ is translated into both a traditional Pig logical plan and an NTGA logical plan; (4) a logical‑to‑physical translator maps NTGA operators to concrete MapReduce jobs; (5) Pig’s job compiler generates an optimized MR workflow. Three new components—rule‑based rewriter, schema‑aware rewriter, and NTGA plan generator—were added to the existing RAPID+ pipeline to support UCQ generation.
Experimental evaluation uses real‑world datasets: UniProt (several terabytes, deep taxonomy hierarchy), Bio2RDF, and Chem2Bio2RDF. A suite of benchmark queries, including complex union queries with over 200 UNION branches, is executed. RAPID+ achieves speed‑ups ranging from 10× to 30× compared with a baseline relational‑style implementation that processes each star‑pattern separately. Moreover, intermediate data volume is reduced by roughly 70 %, alleviating network and disk I/O bottlenecks. In cases where the baseline timed out, RAPID+ completed successfully, demonstrating robustness to query complexity.
The paper’s contributions are threefold: (i) definition of NTGA operators that enable concise representation and efficient parallel execution of high‑width UCQs; (ii) integration of NTGA into a MapReduce‑based query engine (RAPID+), including a complete pipeline from SPARQL parsing to MR job compilation; (iii) thorough empirical validation showing substantial performance gains on large biomedical RDF warehouses. Limitations include increased memory pressure during group formation and potential overhead for extremely sparse property combinations. Future work proposes adaptive group sizing, cost‑based join ordering, and porting the approach to in‑memory distributed frameworks such as Apache Spark.
In summary, the study presents a practical and scalable solution for ontology‑aware query processing on massive life‑science datasets, demonstrating that careful algebraic restructuring (via NTGA) combined with cloud‑scale MapReduce can overcome the bottlenecks inherent in traditional UCQ evaluation.
Comments & Academic Discussion
Loading comments...
Leave a Comment