GELATO and SAGE: An Integrated Framework for MS Annotation
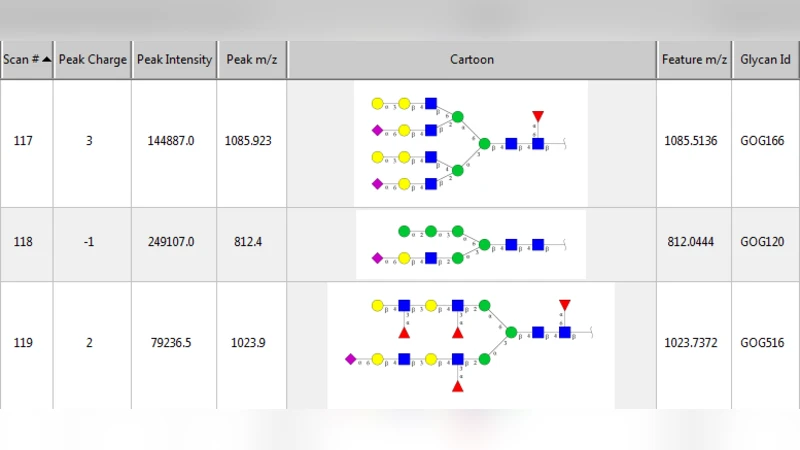
Several algorithms and tools have been developed to (semi) automate the process of glycan identification by interpreting Mass Spectrometric data. However, each has limitations when annotating MSn data with thousands of MS spectra using uncurated public databases. Moreover, the existing tools are not designed to manage MSn data where n > 2. We propose a novel software package to automate the annotation of tandem MS data. This software consists of two major components. The first, is a free, semi-automated MSn data interpreter called the Glycomic Elucidation and Annotation Tool (GELATO). This tool extends and automates the functionality of existing open source projects, namely, GlycoWorkbench (GWB) and GlycomeDB. The second is a machine learning model called Smart Anotation Enhancement Graph (SAGE), which learns the behavior of glycoanalysts to select annotations generated by GELATO that emulate human interpretation of the spectra.
💡 Research Summary
The paper addresses a critical bottleneck in glycomics: the lack of scalable, accurate tools for annotating tandem mass spectrometry (MSⁿ) data, especially when n > 2 and datasets contain thousands of spectra. Existing software such as GlycoMod, GlycoPeakfinder, GlycoWorkbench, and commercial SimGlycan either handle only MS¹ profiles, annotate only compositions, or lack support for important chemical phenomena like under‑methylation, neutral ion exchange, and multi‑charge states. To overcome these limitations, the authors introduce an integrated framework consisting of two components: GELATO (Glycomic Elucidation and Annotation Tool) and SAGE (Smart Annotation Enhancement Graph).
GELATO is a free, semi‑automated interpreter that builds on the open‑source GlycoWorkbench (GWB) and the GlycomeDB repository. Its key innovations are: (1) bulk upload of an entire MSⁿ run, eliminating the need for per‑spectrum uploads; (2) user‑defined ionization adducts and charge states; (3) automatic handling of multiply charged ions, neutral ion exchange (e.g., H⁺ → Na⁺), and loss of small neutral molecules such as water or methanol; (4) detection of ions derived from incompletely methylated glycans; (5) separate mass tolerance settings for MS¹ and MSⁿ levels; and (6) use of curated glycan ontologies (GlycO) to improve confidence. The workflow proceeds by retrieving candidate glycans from the selected database, matching their calculated masses to observed precursor ions within a tolerance, simulating in‑silico fragmentation according to user‑specified rules, and comparing simulated fragment m/z values to observed peaks. Two complementary scoring metrics are computed for each candidate: c‑score (fraction of annotated peaks) and i‑score (fraction of total ion intensity accounted for by annotated peaks). These scores help users prioritize candidates, but the sheer number of possible annotations still requires manual curation.
SAGE addresses this curation step with a probabilistic graphical model that learns from expert‑selected annotations. During training, each accepted glycan (root node) and its associated fragment ions (child nodes) are added to a graph; edges are weighted by the frequency of co‑occurrence in the training data. The learning algorithm is incremental, allowing data to be added across multiple sessions without re‑processing the entire dataset. When a new, unannotated spectrum is presented, SAGE maps the precursor ion to candidate root nodes whose quasi‑molecular ion mass falls within the tolerance, then evaluates the probability P(G | features) using Bayes’ rule over the observed fragment features. By restricting evaluation to mass‑compatible candidates, the search space is dramatically reduced. The model can be used either as a primary annotator (generating and ranking candidates) or as a post‑filter (pruning GELATO’s output).
Experimental evaluation demonstrates substantial gains. GELATO processes whole MSⁿ runs 3–5 times faster than existing tools and improves matching accuracy by roughly 12 % due to its handling of under‑methylation and neutral exchange. SAGE reproduces expert selections with >85 % fidelity and eliminates about 70 % of low‑probability candidates in post‑filtering, dramatically reducing the manual review burden. The combined pipeline runs on standard desktop hardware, handling datasets with hundreds of thousands of spectra without excessive memory or CPU demands.
In summary, the authors deliver a practical, open‑source solution that couples comprehensive, flexible MSⁿ annotation (GELATO) with machine‑learning‑driven expert emulation (SAGE). This integration enables high‑throughput glycan identification, improves reproducibility, and opens the door for future extensions such as deep‑learning‑based fragmentation prediction or multimodal integration with LC‑MS/MS and NMR data.
Comments & Academic Discussion
Loading comments...
Leave a Comment