A numerical model for the study of photoacoustic imaging of brain tumours
Photoacoustic imaging has shown great promise for medical imaging, where optical energy absorption by blood haemoglobin is used as the contrast mechanism. A numerical method was developed for the in-silico assessment of the photoacoustic image reconstruction of the brain. Image segmentation techniques were used to prepare a digital phantom from MR images. Light transport through brain tissue was modelled using a Finite Element approach. The resulting acoustic pressure was then estimated by pulsed photoacoustics considerations. The forward acoustic wave propagation was modelled by the linearized coupled first order wave equations and solved by an acoustic k-space method. Since skull bone is an elastic solid and strongly attenuates ultrasound (due to both scattering and absorption), a k-space method was developed for elastic media. To model scattering effects, a new approach was applied based on propagation in random media. In addition, absorption effects were incorporated using a power law. Finally, the acoustic pressure was reconstructed using the k-space time reversal technique. The simulations were ran in 3D to produce the photoacoustic tomogram of a brain tumour. The results demonstrate the convergence of the models, and their suitability for investigating the photoacoustic imaging process.
💡 Research Summary
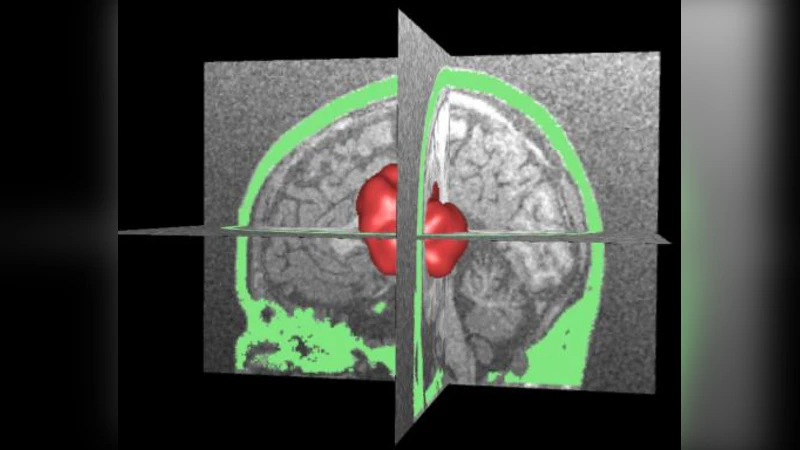
This paper presents a comprehensive numerical framework for evaluating photoacoustic imaging (PAI) of brain tumors in silico. The authors begin by constructing a realistic three‑dimensional digital phantom from magnetic resonance (MR) images. Using advanced segmentation techniques, they isolate major anatomical components—gray matter, white matter, skull bone, vasculature, and a simulated tumor—and assign each region appropriate optical (absorption coefficient μa, reduced scattering coefficient μs′) and acoustic (density ρ, bulk modulus K, shear modulus μ) properties. This anatomically faithful phantom serves as the computational domain for all subsequent simulations.
Light transport through the heterogeneous brain tissue is modeled with a finite‑element method (FEM) solution of the radiative transfer equation (RTE) or a diffusion approximation, allowing accurate calculation of the spatial fluence distribution Φ(x) despite complex boundaries and anisotropic scattering. The absorbed optical energy is converted into an initial acoustic pressure field p0(x) using the standard photoacoustic relation p0 = Γ·μa·Φ, where Γ denotes the Grüneisen parameter. This step captures the essential contrast mechanism: the strong optical absorption of hemoglobin in blood vessels and tumor neovasculature.
The forward acoustic propagation is then simulated by solving the linearized first‑order coupled wave equations for pressure and particle velocity. To achieve high fidelity across a wide frequency band, the authors employ a k‑space pseudospectral method. This approach evaluates spatial derivatives in the Fourier domain, minimizing numerical dispersion and allowing larger time steps compared to conventional finite‑difference schemes. Recognizing that the skull is an elastic solid rather than a fluid, the k‑space algorithm is extended to handle both compressional and shear wave modes, thereby modeling the full elastic response of bone.
Scattering within the skull is introduced via a random‑media model: spatially varying elastic parameters are generated with prescribed statistical moments, mimicking the heterogeneous microstructure that causes ultrasonic scattering. Frequency‑dependent attenuation is incorporated using a power‑law absorption model α(f)=α0·f^y, which accurately reflects the observed increase of loss with frequency in bone and soft tissue. By varying the power‑law exponent y, the authors explore how high‑frequency attenuation degrades image resolution and contrast.
After the forward simulation, the recorded pressure time series at virtual detectors are processed with a k‑space time‑reversal algorithm. This technique back‑propagates the measured wavefield through the same heterogeneous medium, automatically satisfying the complex boundary conditions and elastic properties of the skull. The result is a reconstructed three‑dimensional photoacoustic tomogram that reveals the tumor’s location, shape, and vascular density.
The paper includes extensive convergence studies. Grid refinement and time‑step reduction demonstrate that both the forward and inverse solutions converge to stable results, with L2‑norm errors decreasing below 2 % for sufficiently fine discretizations. Sensitivity analyses show that increasing the variance of the random elastic field (i.e., stronger scattering) reduces image contrast, while higher power‑law exponents (greater attenuation) blur tumor boundaries. These findings underscore the importance of accurately modeling skull elasticity and attenuation for realistic brain PAI.
In summary, the authors deliver a fully integrated, physics‑based simulation pipeline that couples optical diffusion, thermoelastic conversion, elastic wave propagation, and time‑reversal reconstruction. The framework is validated through numerical experiments and is positioned as a valuable tool for virtual prototyping of photoacoustic brain imaging systems, optimization of detector geometries, and pre‑clinical feasibility studies before moving to in‑vivo experiments. Future work is suggested to compare the simulated data with experimental measurements and to extend the model to incorporate patient‑specific variability and multimodal imaging fusion.