Using complex networks towards information retrieval and diagnostics in multidimensional imaging
We present a fresh and broad yet simple approach towards information retrieval in general and diagnostics in particular by applying the theory of complex networks on multidimensional, dynamic images. We demonstrate a successful use of our method with the time series generated from high content thermal imaging videos of patients suffering from the aqueous deficient dry eye (ADDE) disease. Remarkably, network analyses of thermal imaging time series of contact lens users and patients upon whom Laser-Assisted in situ Keratomileusis (Lasik) surgery has been conducted, exhibit pronounced similarity with results obtained from ADDE patients. We also propose a general framework for the transformation of multidimensional images to networks for futuristic biometry. Our approach is general and scalable to other fluctuation-based devices where network parameters derived from fluctuations, act as effective discriminators and diagnostic markers.
💡 Research Summary
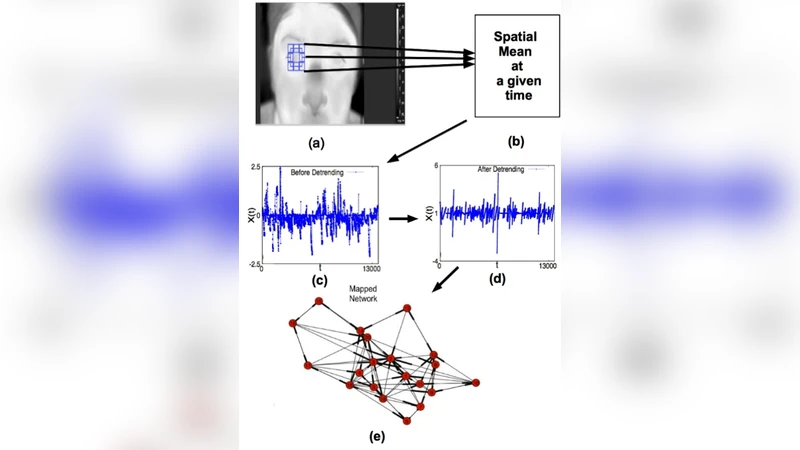
The paper introduces a novel, three‑stage framework that converts multidimensional video data—specifically infrared thermal recordings of the human eye—into a complex‑network representation, and then uses network‑based metrics for information retrieval and diagnostic discrimination. First, each video is reduced to a one‑dimensional time series by extracting the mean temperature of a cropped region of interest (ROI) for every frame. The authors compare this simple averaging approach with principal component analysis (PCA) and demonstrate that the mean‑pixel method retains comparable classification power while being far less computationally demanding, making it suitable for embedded hardware.
Second, each individual time series is detrended to remove a linear decay caused by tear evaporation, normalized, and then pooled across subjects within the same experimental group (healthy controls, aqueous‑deficient dry‑eye patients, contact‑lens wearers with and without lenses, and pre‑/post‑LASIK patients). This pooling yields eight long composite series, denoted {HE}, {DE}, {HC}, {DC}, {CL}, {CN}, {LB}, and {LA}.
Third, the pooled series are mapped onto directed graphs using a variant of the visibility‑graph / transition‑probability method: each time point becomes a node, and directed edges are created according to the relative magnitude and temporal ordering of successive values. The resulting networks are characterized by a suite of topological descriptors—average degree, clustering coefficient, average shortest‑path length, modularity, entropy, and asymmetry measures.
The authors then evaluate whether these descriptors can serve as discriminative biomarkers. Their analysis shows that dry‑eye patients exhibit lower average degree, reduced clustering, and a loss of inter‑ocular temperature correlation compared with healthy eyes. Contact‑lens wear induces similar but less pronounced network alterations, while LASIK surgery leads to a post‑operative decrease in temperature fluctuation and an increase in clustering, reflecting a more stable ocular surface. Importantly, the network‑based features outperform conventional spatial image‑processing metrics, highlighting the value of temporal dynamics that are otherwise invisible in static frames.
Beyond ophthalmology, the authors argue that the pipeline is modality‑agnostic: any sensor that produces a fluctuating multidimensional signal (e.g., ultrasound, functional MRI, optical coherence tomography) can be treated analogously. The method’s low computational load, reliance on simple arithmetic operations, and avoidance of costly dimensionality‑reduction steps make it attractive for real‑time, portable diagnostic devices.
Limitations are acknowledged. Pooling across subjects may mask individual variability, potentially reducing patient‑specific diagnostic precision. The exact graph‑construction rules are not fully formalized, which could hinder reproducibility. Moreover, extending the approach to other imaging modalities will require careful tuning of preprocessing and mapping parameters.
In summary, the study demonstrates that converting biomedical video streams into complex networks provides a powerful, scalable, and hardware‑friendly avenue for information retrieval and disease classification. By exploiting the hidden temporal structure of thermal fluctuations, the authors open a new pathway for non‑invasive diagnostics and biometric applications, with broad implications for future multimodal biomedical imaging research.
Comments & Academic Discussion
Loading comments...
Leave a Comment