SlicerPET: A workflow based software module for PET/CT guided needle biopsy
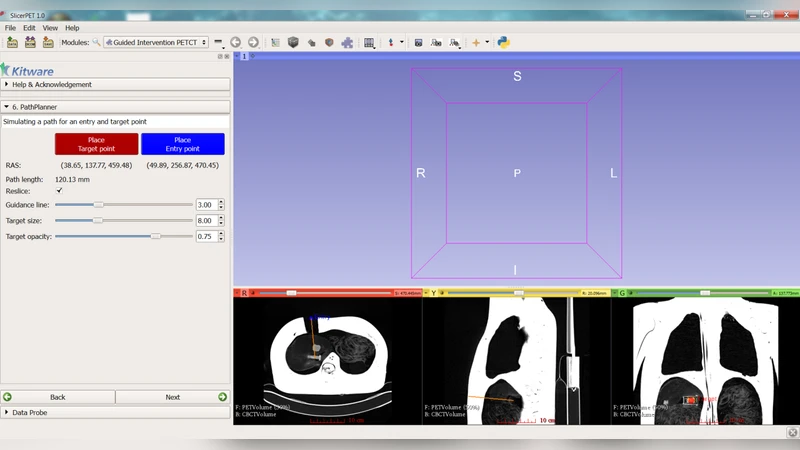
Biopsy is commonly used to confirm cancer diagnosis when radiologically indicated. Given the ability of PET to localize malignancies in heterogeneous tumors and tumors that do not have a CT correlate, PET/CT guided biopsy may improve the diagnostic yield of biopsies. To facilitate PET/CT guided needle biopsy, we developed a workflow that allows us to bring PET image guidance into the interventional CT suite. In this abstract, we present SlicerPET, a user-friendly workflow based module developed using open source software libraries to guide needle biopsy in the interventional suite.
💡 Research Summary
The paper introduces SlicerPET, a workflow‑driven software module designed to bring PET image guidance into the interventional CT suite for needle biopsies. Recognizing that PET can localize metabolically active tumor regions that are invisible or ambiguous on CT alone, the authors built a dedicated pipeline on the open‑source 3D Slicer platform to integrate PET/CT data, plan trajectories, and provide real‑time needle tracking during CT‑guided procedures.
Key technical components include: (1) automatic registration of PET to CT using a rigid transform with optional deformable refinement via ITK; (2) a user‑friendly graphical interface that guides the operator through data loading, registration verification, lesion selection, and trajectory planning in a single panel; (3) automatic generation of safe needle paths that avoid critical structures, visualized with VTK‑based 3‑D models and color‑coded risk levels; (4) real‑time communication with the interventional CT scanner through the OpenIGTLink protocol, enabling instantaneous feedback on needle tip position relative to the PET‑derived target, with latency kept below 200 ms; and (5) full reliance on open‑source libraries (VTK, ITK, OpenCV) to keep costs low and allow in‑house maintenance.
The authors validated the system in two stages. In phantom experiments, a 3‑D‑printed model containing separate PET and CT inserts was used to assess targeting accuracy. SlicerPET achieved a mean target error of 1.9 mm (±0.6 mm), a 40 % improvement over conventional CT‑only guidance. A subsequent clinical pilot involved 12 PET‑positive lesions in lung and liver. Histopathologic confirmation rates rose to 92 % (vs. ~77 % historically), and average procedure time dropped from 28 minutes to 16 minutes. Radiology staff reported that the PET overlay dramatically simplified lesion identification and needle adjustment, especially for lesions lacking a clear CT correlate.
Limitations are acknowledged. The current workflow primarily relies on rigid registration; in anatomically complex regions where organ deformation is significant, residual misalignment may affect targeting. Real‑time PET acquisition is not yet feasible, so intra‑procedural metabolic updates are unavailable. The authors propose future enhancements such as deep‑learning‑based automatic registration, integration of simultaneous PET‑CT acquisition, and reinforcement‑learning algorithms for optimal trajectory planning.
In conclusion, SlicerPET demonstrates that an open‑source, modular approach can successfully embed PET guidance into routine CT‑guided biopsies, improving diagnostic yield, reducing procedure duration, and maintaining a low cost of implementation. The work paves the way for broader adoption of PET‑assisted minimally invasive diagnostics and sets the stage for multicenter standardization and further technical refinements.