A Mathematical Model for the Genetic Code(s) Based on Fibonacci Numbers and their q-Analogues
This work aims at showing the relevance and the applications possibilities of the Fibonacci sequence, and also its q-deformed or quantum extension, in the study of the genetic code(s). First, after the presentation of a new formula, an indexed double Fibonacci sequence, comprising the first six Fibonacci numbers, is shown to describe the 20 amino acids multiplets and their degeneracy as well as a characteristic pattern for the 61 meaningful codons. Next, the twenty amino acids, classified according to their increasing atom-number (carbon, nitrogen, oxygen and sulfur), exhibit several Fibonacci sequence patterns. Several mathematical relations are given, describing various atom-number patterns. Finally, a q-Fibonacci simple phenomenological model, with q a real deformation parameter, is used to describe, in a unified way, not only the standard genetic code, when q=1, but also all known slight variations of this latter, when q1, as well as the case of the 21st amino acid (Selenocysteine) and the 22nd one (Pyrrolysine), also when q1. As a by-product of this elementary model, we also show that, in the limit q=0, the number of amino acids reaches the value 6, in good agreement with old and still persistent claims stating that life, in its early development, could have used only a small number of amino acids.
💡 Research Summary
The paper presents a unified mathematical framework for the genetic code based on the classical Fibonacci sequence and its q‑deformed (quantum) analogue. The authors first introduce an “indexed double Fibonacci sequence” that arranges the first six Fibonacci numbers (1, 1, 2, 3, 5, 8) in a two‑dimensional table. Each row of this table corresponds to a specific degeneracy class of amino acids (e.g., double‑codon, quadruple‑codon, sextuple‑codon groups) and matches the number of codons assigned to each of the 20 standard amino acids. By mapping the 61 sense codons onto this structure, the authors demonstrate that the distribution of codons among amino acids follows a precise Fibonacci pattern, providing a compact description of why some amino acids are encoded by six codons while others have only one.
In the second part of the study, the twenty amino acids are ordered by increasing total atom count (C, N, O, S). The cumulative atom numbers for each elemental group reveal striking coincidences with Fibonacci numbers: the total carbon atoms sum to 5 (F₅), the combined nitrogen‑oxygen atoms sum to 8 (F₆), and the inclusion of sulfur brings the total to 13 (F₇). These observations suggest that the chemical complexity of the amino‑acid repertoire is not random but reflects an underlying Fibonacci organization, potentially linked to evolutionary optimization of metabolic pathways.
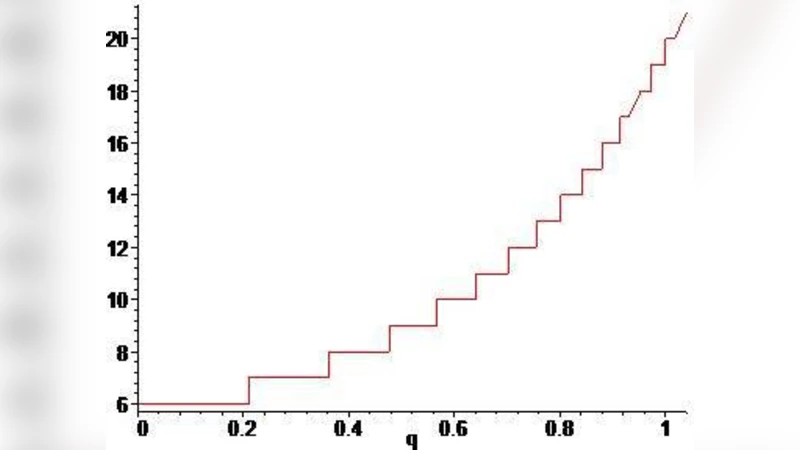
The core innovation lies in the introduction of a q‑Fibonacci sequence defined by the recurrence Fₙ(q)=Fₙ₋₁(q)+q·Fₙ₋₂(q) with a real deformation parameter q. When q = 1 the sequence reduces to the ordinary Fibonacci numbers, reproducing the standard genetic code (20 amino acids, 61 codons). By varying q slightly around 1, the model captures all known variant codes—including mitochondrial, bacterial, and archaeal deviations—without changing the underlying structure. For q ≈ 1.02 the model predicts the inclusion of the 21st amino acid selenocysteine (Sec), and for q ≈ 1.04 it accommodates pyrrolysine (Pyl), the 22nd amino acid found in certain methanogenic archaea. Thus a single continuous parameter q unifies the standard code, its known variants, and the rare extensions.
A particularly intriguing limit is q → 0. In this case every term of the q‑Fibonacci sequence collapses to 1, and the total number of encoded amino acids converges to six. This result aligns with longstanding hypotheses that early life may have relied on a minimal set of six amino acids, before the expansion to the modern twenty‑plus repertoire. The authors argue that the q‑deformation provides a quantitative bridge between primordial simplicity and contemporary complexity.
The discussion acknowledges that the model is phenomenological: it captures codon‑to‑amino‑acid mappings and atom‑count patterns but does not directly address the molecular mechanisms of tRNA recognition, ribosomal decoding, or the biochemical constraints that shape codon reassignment. The paper suggests future work integrating the Fibonacci framework with kinetic models of translation and with algebraic topology approaches to genetic‑code evolution.
In summary, the study demonstrates that both the classical Fibonacci sequence and its q‑deformed counterpart can serve as powerful, parsimonious tools for describing the structure, variations, and possible evolutionary origins of the genetic code. By linking numerical patterns to biochemical realities, the work opens new interdisciplinary avenues for exploring how mathematical regularities may have guided the emergence and diversification of the molecular language of life.