Stochastic modeling of gene expression, protein modification, and polymerization
Many fundamental cellular processes involve small numbers of molecules. When numbers are small, fluctuations dominate, and stochastic models, which account for these fluctuations, are required. In this chapter, we describe minimal stochastic models of three fundamental cellular processes: gene expression, protein modification, and polymerization. We introduce key analytic tools for solving each model, including the generating function, eigenfunction expansion, and operator methods, and we discuss how these tools are extended to more complicated models. These analytic tools provide an elegant, efficient, and often insightful alternative to stochastic simulation.
💡 Research Summary
**
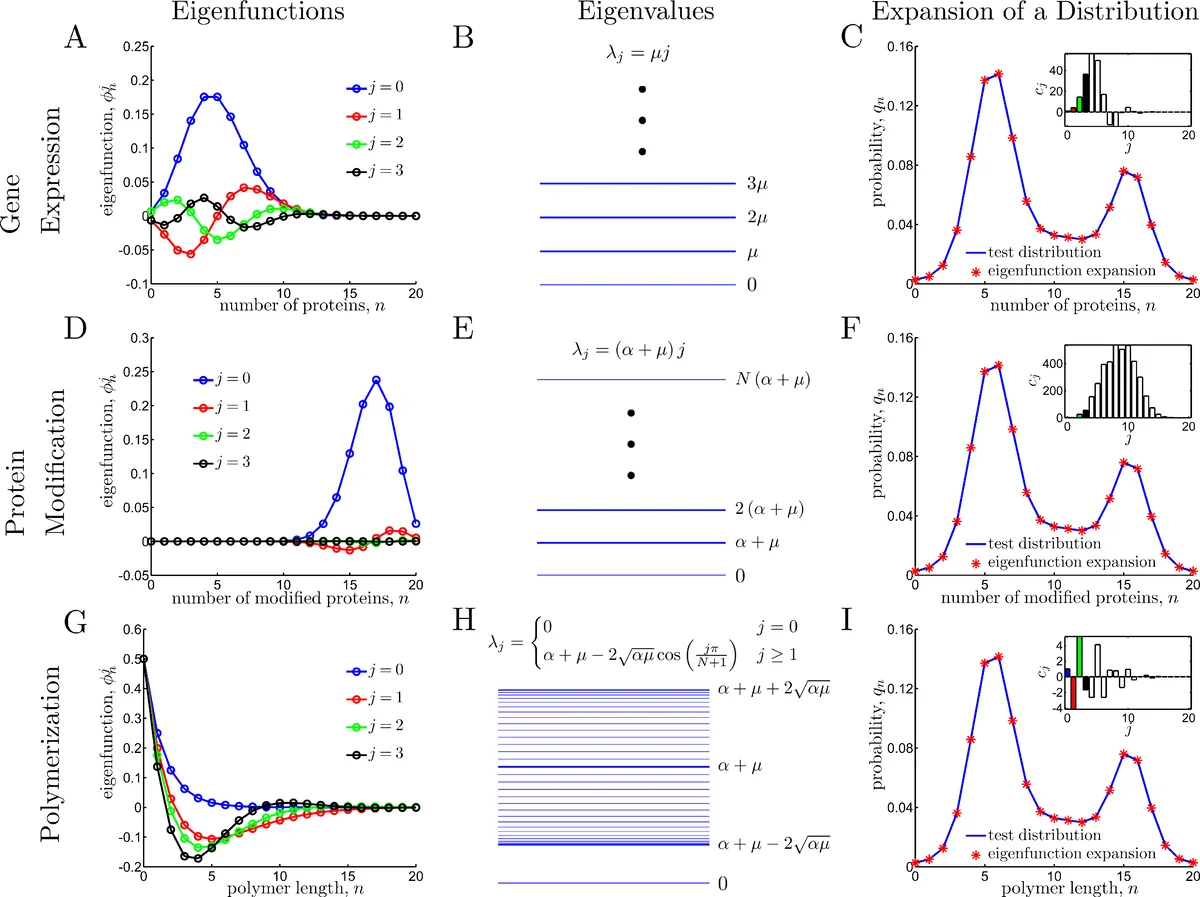
This chapter presents exact stochastic analyses of three fundamental cellular processes—gene expression, protein modification, and polymerization—using three complementary analytical tools: generating functions, eigenfunction expansions, and operator methods. Each process is reduced to a minimal birth‑death–type model that captures the essential stochastic dynamics while remaining analytically tractable.
Gene expression is modeled as a simple birth‑death process with constant protein synthesis rate α and degradation/dilution rate μ. The master equation for the probability pₙ(t) of having n proteins leads, in steady state, to a Poisson distribution pₙ = e⁻ᵞ γⁿ/n! with γ = α/μ. Introducing the generating function G(z)=∑ₙpₙzⁿ converts the infinite set of coupled ODEs into a single PDE: ∂ₜG = –(z–1)(μ∂_z – α)G. The steady‑state solution is G(z)=e⁻ᵞ e^{γz}, and the full time‑dependent solution is obtained either by the method of characteristics or by a change of variables, yielding G(z,t)=F
Comments & Academic Discussion
Loading comments...
Leave a Comment