The XL-mHG Test For Enrichment: A Technical Report
The minimum hypergeometric test (mHG) is a powerful nonparametric hypothesis test to detect enrichment in ranked binary lists. Here, I provide a detailed review of its definition, as well as the algorithms used in its implementation, which enable the efficient computation of an exact p-value. I then introduce a generalization of the mHG, termed XL-mHG, which provides additional control over the type of enrichment tested, and describe the precise algorithmic modifications necessary to compute its test statistic and p-value. The XL-mHG algorithm is a building block of GO-PCA, a recently proposed method for the exploratory analysis of gene expression data using prior knowledge.
💡 Research Summary
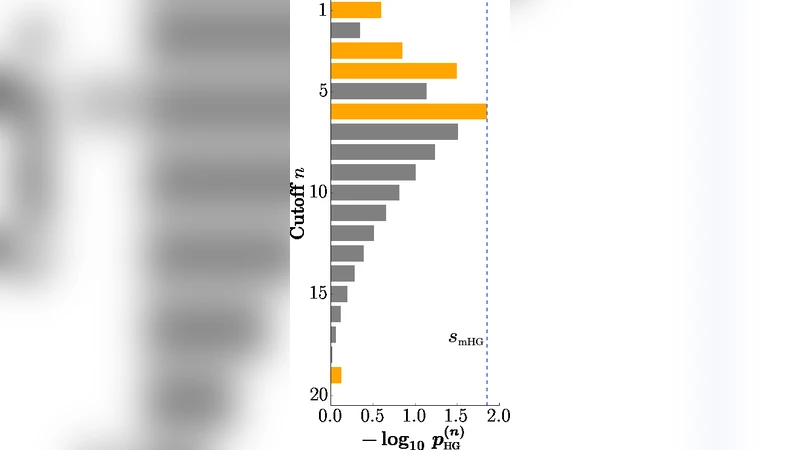
The paper provides a thorough exposition of the minimum hypergeometric (mHG) test, a non‑parametric method for detecting enrichment in ranked binary lists, and introduces its extension, the XL‑mHG test. The authors first formalize a ranked binary list as a vector v of length N containing only 0s and 1s, with K ones and W zeros (N = K + W). A naïve enrichment test would fix a cutoff n, count the number of ones k(n) in the top n positions, and compute a hypergeometric p‑value p_HG(n). This approach suffers from a severe bias because the choice of n is arbitrary and can dramatically affect significance.
The mHG test eliminates the need for a pre‑specified cutoff by evaluating p_HG(n) for every possible n (1 ≤ n ≤ N) and taking the minimum value s_mHG = min_n p_HG(n) as the test statistic. Although this introduces a multiple‑testing problem, the authors show that an exact p‑value p_mHG can be obtained efficiently via dynamic programming that counts admissible paths through the (n, k) space. They also present the Lipson bound p_mHG ≤ K·s_mHG, which provides a fast upper bound.
Despite its strengths, mHG has two practical limitations: (1) when K is small, the minimum p‑value can be overly optimistic, and (2) the test implicitly scans the entire list, which may dilute power when the true signal resides only near the top. To address these issues, the paper proposes the XL‑mHG test, adding two user‑controlled parameters: X (the minimum number of ones required for a candidate enrichment) and L (the maximum rank to be examined). By restricting the search to positions n ≤ L and ignoring cutoffs where k < X, XL‑mHG reduces false positives caused by tiny sub‑sets and focuses the analysis on the biologically relevant region of the list.
Algorithmically, XL‑mHG modifies the mHG dynamic‑programming tables: rows corresponding to k < X and columns corresponding to n > L are masked out before the minimization step. The resulting test statistic s_XL‑mHG and exact p‑value p_XL‑mHG are computed with the same O(N·K) complexity as the original mHG, but often with far fewer operations because many (n, k) pairs are excluded. The authors also define enrichment strength scores e_mHG and e_XL‑mHG, which combine the test statistic and its p‑value on a log scale and normalize by K or X, providing an intuitive measure of effect size.
The implementation is released as open‑source Cython code (https://github.com/flo-compbio/xlmhg), leveraging SciPy’s hypergeometric functions for probability calculations and optimized memory usage. The paper demonstrates the utility of XL‑mHG within GO‑PCA, a knowledge‑driven unsupervised analysis framework for gene‑expression data. In GO‑PCA, each Gene Ontology term is represented as a binary vector of genes; XL‑mHG quantifies the enrichment of each term in the ranked expression list, enabling robust dimensionality reduction and biologically meaningful interpretation.
In summary, the work delivers a complete theoretical foundation, efficient exact algorithms, and practical extensions for enrichment testing in large‑scale ranked binary data. By introducing controllable parameters X and L, XL‑mHG retains the non‑parametric power of mHG while offering greater specificity and interpretability for real‑world applications such as functional genomics.
Comments & Academic Discussion
Loading comments...
Leave a Comment