Control and controllability of nonlinear dynamical networks: a geometrical approach
In spite of the recent interest and advances in linear controllability of complex networks, controlling nonlinear network dynamics remains to be an outstanding problem. We develop an experimentally feasible control framework for nonlinear dynamical networks that exhibit multistability (multiple coexisting final states or attractors), which are representative of, e.g., gene regulatory networks (GRNs). The control objective is to apply parameter perturbation to drive the system from one attractor to another, assuming that the former is undesired and the latter is desired. To make our framework practically useful, we consider RESTRICTED parameter perturbation by imposing the following two constraints: (a) it must be experimentally realizable and (b) it is applied only temporarily. We introduce the concept of ATTRACTOR NETWORK, in which the nodes are the distinct attractors of the system, and there is a directional link from one attractor to another if the system can be driven from the former to the latter using restricted control perturbation. Introduction of the attractor network allows us to formulate a controllability framework for nonlinear dynamical networks: a network is more controllable if the underlying attractor network is more strongly connected, which can be quantified. We demonstrate our control framework using examples from various models of experimental GRNs. A finding is that, due to nonlinearity, noise can counter-intuitively facilitate control of the network dynamics.
💡 Research Summary
The paper tackles the long‑standing challenge of controlling nonlinear dynamical networks, especially those that exhibit multistability—a hallmark of many biological systems such as gene regulatory networks (GRNs). While linear controllability theory (based on Kalman’s rank condition) has yielded powerful graph‑theoretic tools for complex networks, it cannot be applied when node dynamics are intrinsically nonlinear. To bridge this gap, the authors introduce a novel, experimentally grounded framework that relies on two key ideas: (1) the construction of an “attractor network” and (2) the use of temporary, bounded parameter perturbations to drive the system from an undesired attractor to a desired one.
Attractor network concept.
Every stable attractor (fixed point, limit cycle, etc.) of the original high‑dimensional system is represented as a node in a coarse‑grained graph. A directed edge from attractor A to attractor B exists if, by applying a feasible, time‑limited change to one (or a few) system parameters, the dynamics can be forced to leave A’s basin of attraction and eventually settle in B’s basin after the perturbation is removed. Multiple edges may connect the same pair of attractors, each corresponding to a different controllable parameter. This graph abstracts the full phase space while preserving precisely the information needed for the control task.
Control mechanism.
The authors focus on bifurcation‑based transitions. By temporarily shifting a control parameter µ from its nominal value µ₀ to a perturbed value µ₁, the system is moved close to a bifurcation point (typically a saddle‑node). At µ₁ the original attractor disappears, and an intermediate attractor ¯x* appears. The trajectory then follows the flow toward ¯x* while the perturbation is active. When the perturbation is withdrawn (µ returns to µ₀), the system finds itself inside the basin of the target attractor x*_f and converges to it. If a single parameter cannot create a viable path, the framework allows simultaneous perturbations of several parameters—conceptually equivalent to applying a drug cocktail in a biological context.
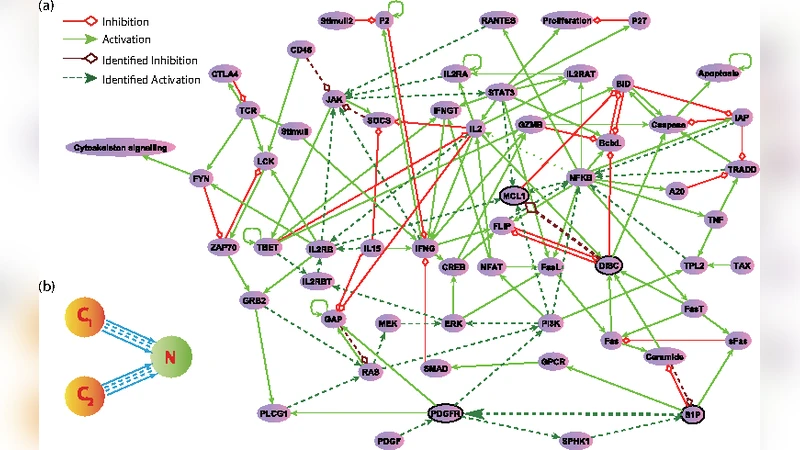
Demonstration on a realistic GRN.
The methodology is illustrated on a 60‑node, 142‑edge T‑cell survival signaling network (T‑LGL leukemia model). Boolean update rules are translated into continuous ordinary differential equations, and the system is shown to possess three stable fixed points: two cancerous states (C₁, C₂) and one normal state (N). Exhaustive testing of all 48 adjustable edges reveals numerous viable control edges that can drive either cancerous attractor to the normal one. Control pulses are modeled as rectangular pulses of duration τ and amplitude Δµ. A systematic analysis uncovers a power‑law relationship τₘ ∝ (Δe)^β between the minimal control time τₘ and the excess parameter change Δe = µ_c – µ_n (β ≈ 0.4), indicating that larger perturbations require shorter intervention times.
Counter‑intuitive role of noise.
An important finding is that stochastic fluctuations can facilitate control. Moderate noise pushes the system toward bifurcation thresholds, effectively lowering the required Δµ or τ for successful transition. This “noise‑facilitated control” parallels stochastic resonance but is framed here as a beneficial element for achieving a prescribed state change rather than merely enhancing signal detection.
Quantifying controllability.
The connectivity of the attractor network provides a natural metric for the controllability of the underlying nonlinear system. Strongly connected components, average shortest‑path lengths, and edge multiplicities quantify how easily any attractor can be reached from any other. A densely connected attractor network implies high controllability, whereas a sparsely linked one signals the need for larger or more complex perturbations.
Implications and applications.
Because the framework explicitly respects experimental constraints—only temporary, physically realizable parameter changes are allowed—it is directly applicable to synthetic biology, drug‑combination therapy design, and cell‑fate engineering. For instance, one can pre‑compute the minimal set of drug targets and dosing durations needed to reprogram cancer cells into a normal phenotype, while keeping interventions short to minimize side effects.
In summary, the paper provides a concrete, geometrically inspired approach to controlling multistable nonlinear networks. By abstracting the phase space into an attractor network and exploiting bifurcation‑induced transitions, it offers a scalable, experimentally viable pathway to steer complex biological systems toward desired functional states, and it highlights the surprising constructive role that noise can play in such control tasks.
Comments & Academic Discussion
Loading comments...
Leave a Comment