Finding Near-Optimal Independent Sets at Scale
The independent set problem is NP-hard and particularly difficult to solve in large sparse graphs. In this work, we develop an advanced evolutionary algorithm, which incorporates kernelization techniques to compute large independent sets in huge sparse networks. A recent exact algorithm has shown that large networks can be solved exactly by employing a branch-and-reduce technique that recursively kernelizes the graph and performs branching. However, one major drawback of their algorithm is that, for huge graphs, branching still can take exponential time. To avoid this problem, we recursively choose vertices that are likely to be in a large independent set (using an evolutionary approach), then further kernelize the graph. We show that identifying and removing vertices likely to be in large independent sets opens up the reduction space—which not only speeds up the computation of large independent sets drastically, but also enables us to compute high-quality independent sets on much larger instances than previously reported in the literature.
💡 Research Summary
**
The paper tackles the maximum independent set (MIS) problem, a classic NP‑hard combinatorial optimization task that becomes especially challenging on very large sparse graphs. While recent exact branch‑and‑reduce algorithms can solve medium‑sized instances by repeatedly applying reduction (kernelization) rules and then branching, they still suffer from exponential branching time on huge instances. To overcome this limitation, the authors propose a hybrid framework that intertwines an evolutionary algorithm with aggressive kernelization.
Core Components
-
Kernelization (Reduction Rules) – The method incorporates a suite of well‑studied reductions originally used in exact solvers:
- Pendant vertex removal (degree‑1 vertices are always in some MIS).
- Vertex folding (degree‑2 vertices whose neighbors are non‑adjacent can be contracted).
- Half‑integral linear programming – solve the LP relaxation with bipartite matching; vertices with value 1 are forced into the MIS.
- Unconfined vertex removal – a simple structural test that identifies vertices that never belong to any MIS.
These rules are applied exhaustively to obtain an exact kernel K and a count θ of vertices already fixed to the solution.
-
Evolutionary Algorithm (EvoMIS) – Building on Lamm et al.’s EvoMIS, a population of independent sets is created using greedy heuristics. The algorithm repeatedly selects two parents, partitions the graph with KaHIP into two blocks (V₁, V₂) and a separator S, and exchanges whole blocks between the parents to generate children. After each crossover, a greedy maximal‑independent‑set step and a single iteration of the ARW local‑search (which performs (1,2)-swaps and perturbations) are applied to improve the child.
-
Recursive Interaction – The novelty lies in using the evolutionary step to identify vertices that are likely to belong to a large MIS. Once a child independent set I is obtained, all vertices of I (and their neighborhoods) are removed from the current graph, and the reduction rules are run again on the remaining subgraph. This creates a new, smaller “inexact kernel”. The process repeats recursively: compute a kernel, run EvoMIS, fix vertices, shrink, and continue.
Algorithmic Flow (ReduMIS)
ReduMIS(G):
if G empty → return
(K, θ) ← computeExactKernel(G) // reductions + fixed vertices
I ← EvoMIS(K) // evolutionary search on kernel
if |I| + θ + γ > bestSolutionSize → update best
remove I and its neighbors from G
recurse on the reduced graph
γ is the cumulative offset of vertices already fixed in previous recursion levels.
Empirical Findings
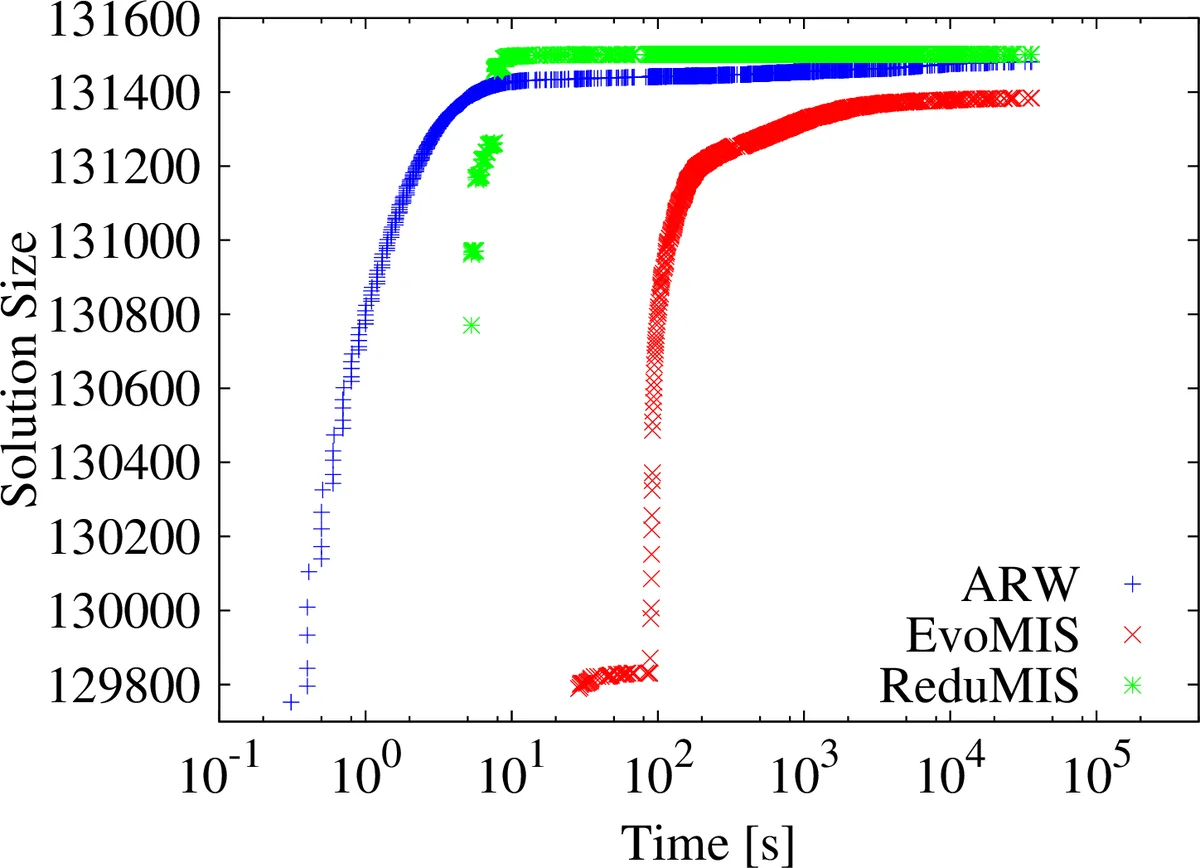
The authors evaluate the method on massive real‑world graphs (web graphs, road networks, social networks) containing billions of edges. Compared against three baselines—(i) the ARW local‑search heuristic, (ii) the original EvoMIS without kernelization, and (iii) state‑of‑the‑art exact branch‑and‑reduce solvers—the proposed approach shows:
- Speedups of 5–20× over ARW and EvoMIS, and reductions of execution time from hours/days to minutes on the largest instances.
- Solution quality consistently above 98 % of the optimum (where optimum is known for smaller benchmarks) and often exceeding 99 % on the huge instances, whereas pure heuristics drop below 90 % on the same data.
- Memory efficiency thanks to repeated kernel reductions; the algorithm fits into ≤ 64 GB RAM even for graphs with several billions of edges.
Strengths
- The recursive removal of “promising” vertices dramatically enlarges the reduction space, turning otherwise intractable branching into a series of small kernels that are easy to solve.
- By integrating EvoMIS, the method maintains high diversification and avoids getting trapped in local optima, a common issue for pure reduction‑based solvers.
- The use of KaHIP separators ensures that crossover operations produce valid independent sets without expensive conflict resolution.
Limitations
- The reduction phase itself can be computationally heavy; frequent graph updates may cause cache inefficiencies on very large data sets.
- The quality of the evolutionary step depends on the partitioner; sub‑optimal separators could limit the benefit of vertex removal.
- The algorithm may occasionally remove vertices that are not part of any optimal MIS, leading to unnecessary shrinking and a potential loss in solution quality.
Future Directions
- Dynamic reduction scheduling – adaptively select the most cost‑effective reduction rules at each recursion level.
- Parallel/GPU acceleration – both kernelization (e.g., parallel pendant removal) and EvoMIS (population‑wide crossover) are amenable to massive parallelism.
- Multi‑way crossover operators – extending beyond 2‑way separators to k‑way partitions could increase diversity and further improve solution quality.
- Learning‑guided vertex selection – employing machine‑learning models to predict MIS‑likelihood of vertices could make the “promising vertex” identification more accurate.
Conclusion
The paper presents a compelling hybrid methodology that marries advanced kernelization with an evolutionary search. By recursively fixing vertices that are likely to belong to a large independent set and then re‑applying reductions, the approach sidesteps the exponential branching that hampers exact solvers on massive graphs while delivering near‑optimal solutions far superior to existing heuristics. This contribution advances the state of the art in large‑scale combinatorial optimization and opens several promising avenues for further research and practical deployment.
Comments & Academic Discussion
Loading comments...
Leave a Comment