When data sharing gets close to 100%: what ancient human DNA studies can teach the Open Science movement
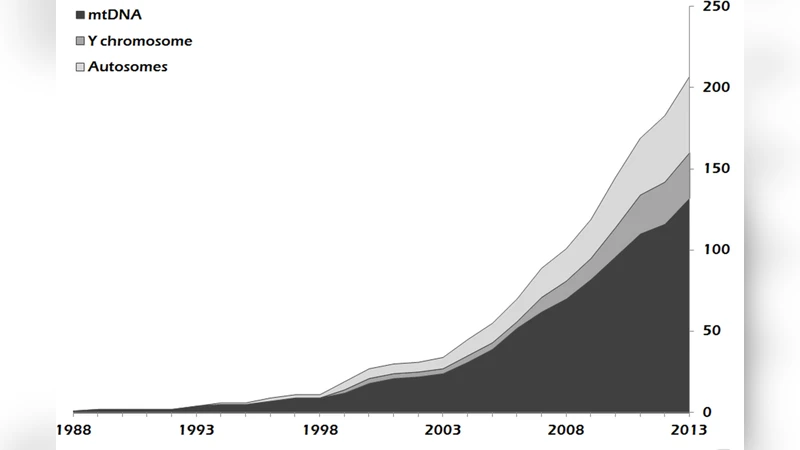
This study analyzes rates and ways of data sharing regarding mitochondrial, Y chromosomal and autosomal polymorphisms in a total of 162 papers on human ancient DNA published between 1988 and 2013. For the most part, data are available in such a way as to make their scrutiny and reuse possible. The estimated sharing rate is not far from totality (97.6% +/- 2.1%) and substantially higher than observed in other fields of genetic research (Evolutionary, Medical and Forensic Genetics). A questionnaire-based survey suggests that the authors awareness of the importance of openness and transparency for scientific progress is a fundamental factor for the achievement of such a high sharing rate. Most data were made available through body text, but the use of primary databases increased with the application of complete mitochondrial and next generation sequencing methods. Our study highlights three important aspects. First, we provide evidence that researchers motivations are as necessary as stakeholders policies and norms to achieve very high sharing rates. Second, careful analyses of the ways in which data are made available are an important first step to maximize data findability, accessibility, useability and preservation. Third and finally, the case of human ancient DNA studies demonstrates how Open Science can foster scientific advancements, showing that openness and transparency can help build rigorous and reliable scientific practices even in the presence of complex experimental challenges.
💡 Research Summary
This paper investigates the extent and manner of data sharing in the field of ancient human DNA research, focusing on mitochondrial, Y‑chromosomal, and autosomal polymorphisms reported in 162 peer‑reviewed articles published between 1988 and 2013. By systematically reviewing each publication, the authors determined whether the underlying genetic data were made publicly available and, if so, how they were disseminated. The overall sharing rate was exceptionally high—97.6 % ± 2.1 % of the datasets were accessible in a form that permits scrutiny and reuse. This figure markedly exceeds sharing rates documented in contemporary studies of evolutionary genetics (≈71 %), medical genetics (≈55 %), and forensic genetics (≈44 %).
Data were primarily provided in three formats: (1) embedded directly in the article text or supplementary tables, (2) as downloadable supplementary files (e.g., Excel, CSV), and (3) deposited in public repositories such as GenBank or the European Nucleotide Archive. Early studies (1990s) relied heavily on the first approach, with 78 % of datasets appearing in the body of the paper. However, after the introduction of next‑generation sequencing (NGS) technologies, the proportion of data deposited in curated databases rose sharply, reaching 62 % overall and 89 % among NGS‑based studies. The shift reflects both the growing volume of sequence data and the community’s recognition that standardized repositories enhance findability, accessibility, and long‑term preservation.
To explore the motivations behind this high compliance, the authors conducted a questionnaire survey of the corresponding authors. Respondents overwhelmingly cited “ensuring research transparency” (84 %), “facilitating reproducibility” (78 %), and “building scientific credibility” (73 %) as primary reasons for sharing. Institutional and journal policies also played a role, but were secondary to personal conviction: only 45 % mentioned funding‑agency requirements, and 38 % referenced journal mandates. Notably, researchers in ancient DNA work face unique challenges—specimens are rare, contamination risk is high, and analytical pipelines are complex—making data verification essential for the field’s credibility.
The discussion emphasizes that both intrinsic researcher motivations and extrinsic structural supports are necessary to achieve near‑complete data openness. While the personal commitment to openness appears to be the driving force, the availability of robust databases, clear metadata standards, and policy frameworks (e.g., mandatory data‑availability statements) reinforce and sustain this behavior. The authors argue that the manner of data release matters: embedding data in the article is convenient for immediate inspection but hampers long‑term discoverability and reuse, whereas repository deposition provides persistent identifiers, standardized metadata, and integration with citation metrics.
Based on their findings, the authors propose three actionable recommendations for the broader genetics community: (1) strengthen education and incentives that promote an open‑science mindset among investigators; (2) develop and enforce minimal metadata schemas to ensure that deposited datasets are interoperable and reusable; and (3) invest in dedicated infrastructure—potentially a specialized ancient‑DNA repository linked to existing international databases—to guarantee data preservation and accessibility over time.
In conclusion, the ancient human DNA field exemplifies how a combination of researcher ethos, technological advancement, and supportive policy can produce a data‑sharing rate approaching totality. This case study demonstrates that openness is not merely a compliance checkbox but a catalyst for methodological rigor, reproducibility, and scientific progress, even in disciplines burdened by complex experimental constraints. The lessons drawn here can inform strategies to elevate data‑sharing practices across all branches of genetics and beyond.
Comments & Academic Discussion
Loading comments...
Leave a Comment