The viscoelastic properties of chromatin and the nucleoplasm revealed by scale-dependent protein mobility
The eukaryotic cell nucleus harbors the DNA genome that is organized in a dynamic chromatin network and embedded in a viscous crowded fluid. This environment directly affects enzymatic reactions and target search processes that access the DNA sequence information. However, its physical properties as a reaction medium are poorly understood. Here, we exploit mobility measurements of differently sized inert green fluorescent tracer proteins to characterize the viscoelastic properties of the nuclear interior of a living human cell. We find that it resembles a viscous fluid on small and large scales, but appears viscoelastic on intermediate scales that change with protein size. Our results are consistent with simulations of diffusion through polymers and suggest that chromatin forms a random obstacle network rather than a self-similar structure with fixed fractal dimension. By calculating how long molecules remember their previous position in dependence on their size, we evaluate how the nuclear environment affects search processes of chromatin targets.
💡 Research Summary
The authors set out to quantify the physical nature of the eukaryotic nucleus, focusing on how the chromatin mesh and the surrounding nucleoplasmic fluid influence the mobility of inert macromolecules. To this end, they engineered five green fluorescent protein (GFP) variants that span a hydrodynamic radius range of roughly 2 nm to 10 nm and expressed them in living HeLa cells. Because these tracers are inert and do not bind DNA, their motion reflects only the mechanical properties of the nuclear interior.
Mobility was measured using two complementary techniques. High‑speed single‑particle tracking (SPT) recorded three‑dimensional trajectories at >1 kHz, allowing calculation of the mean‑square displacement (MSD) as a function of lag time τ. In parallel, fluorescence recovery after photobleaching (FRAP) was performed on defined nuclear regions to obtain effective diffusion coefficients. Both methods yielded consistent MSD curves, confirming that experimental artefacts were minimal.
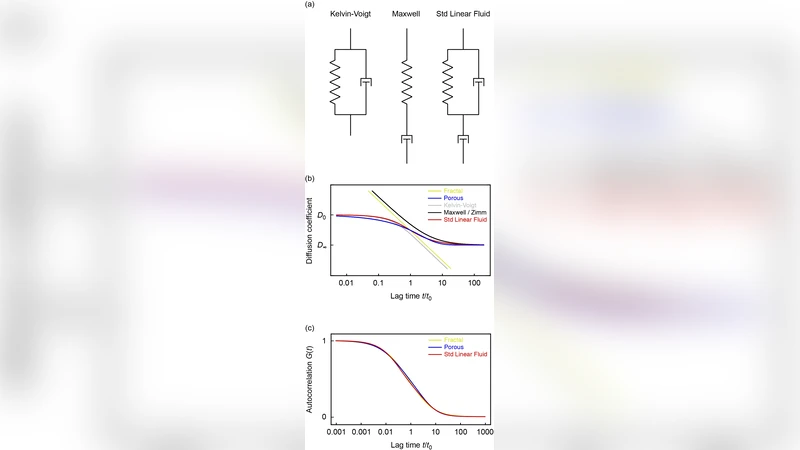
Analysis of the MSD revealed three distinct regimes. At very short lag times (τ ≤ 10 ms) the MSD scales linearly with τ (exponent α≈1), indicating pure viscous diffusion. At intermediate times (10 ms < τ < 1 s) the scaling exponent drops to 0.7–0.9, a hallmark of viscoelastic behavior. Importantly, α decreases systematically with tracer size, showing that larger particles experience a more constrained environment. At long times (τ ≥ 1 s) the exponent returns to ≈1, suggesting that on macroscopic nuclear scales the medium behaves again as a simple viscous fluid.
To translate these observations into mechanical parameters, the complex shear modulus G*(ω)=G′(ω)+iG″(ω) was extracted via the generalized Stokes–Einstein relation. At low frequencies (ω < 10 rad s⁻¹) the loss modulus G″ dominates, confirming a viscous regime. In the intermediate band (10–100 rad s⁻¹) storage and loss moduli intersect, producing a peak in elastic response that marks the viscoelastic window. At higher frequencies the loss modulus regains dominance, reverting to a purely viscous response. This frequency‑dependent crossover mirrors predictions for semi‑dilute polymer networks rather than for a self‑similar fractal scaffold.
To test the structural hypothesis, the authors performed Langevin dynamics simulations of a three‑dimensional random obstacle network. Polymers of varying length and orientation were placed randomly to generate a “random obstacle” (random‑obstacle) matrix, and tracer particles of the same sizes as the experimental GFPs were allowed to diffuse. The simulated MSD and G* curves matched the experimental data closely, reproducing the size‑dependent reduction in α and the G′/G″ crossover. This agreement supports the view that chromatin behaves as a random obstacle mesh with a characteristic mesh size that is probed differently by particles of different dimensions.
Beyond static rheology, the study examined dynamic memory effects by computing the autocorrelation‑based memory kernel K(t)=⟨Δr(0)·Δr(t)⟩/⟨Δr(0)²⟩. Small tracers (≈2 nm) lose correlation within ~20 ms, whereas the largest tracers retain memory for >200 ms. This size‑dependent “memory time” implies that large nuclear complexes—such as transcription factors, polymerases, or CRISPR‑Cas assemblies—experience a temporally correlated environment that can slow target search. By integrating the measured memory kernels into first‑passage‑time models, the authors showed that the average time to locate a specific DNA locus can increase by a factor of 2–5 relative to pure diffusion, depending on particle size and chromatin density.
The discussion contrasts the random‑obstacle picture with earlier fractal models that assume a fixed fractal dimension and scale‑free diffusion. While fractal models predict a universal sub‑diffusive exponent independent of particle size, the present data demonstrate a clear size dependence, undermining the fractal hypothesis for the nuclear interior. Instead, the random‑obstacle framework naturally accounts for the observed crossover from viscous to viscoelastic to viscous behavior and for the particle‑size‑dependent memory.
Finally, the authors consider biological implications. The viscoelastic window (≈10 ms–1 s) coincides with the timescale of many nuclear processes, such as transcription factor binding, nucleosome remodeling, and DNA repair. The enhanced elasticity at intermediate frequencies may act as a mechanical filter, temporarily trapping proteins near chromatin and thereby increasing the probability of productive interactions. Conversely, the eventual return to viscous behavior at long times ensures that macromolecules can still explore the entire nuclear volume over minutes, preserving overall genome accessibility.
In summary, this work provides a comprehensive, multi‑scale characterization of nuclear mechanics in living human cells. By combining size‑controlled inert tracers, high‑resolution tracking, rheological analysis, and polymer simulations, the authors demonstrate that the nucleus behaves as a viscous fluid at very short and very long length scales but exhibits size‑dependent viscoelasticity at intermediate scales. The findings favor a random‑obstacle model of chromatin organization and offer quantitative tools for predicting how nuclear rheology influences biochemical reaction rates, gene regulation, and the efficiency of genome‑editing technologies.