An Iterative Convolutional Neural Network Algorithm Improves Electron Microscopy Image Segmentation
To build the connectomics map of the brain, we developed a new algorithm that can automatically refine the Membrane Detection Probability Maps (MDPM) generated to perform automatic segmentation of electron microscopy (EM) images. To achieve this, we executed supervised training of a convolutional neural network to recover the removed center pixel label of patches sampled from a MDPM. MDPM can be generated from other machine learning based algorithms recognizing whether a pixel in an image corresponds to the cell membrane. By iteratively applying this network over MDPM for multiple rounds, we were able to significantly improve membrane segmentation results.
💡 Research Summary
The paper addresses a central challenge in connectomics: the reliable automatic segmentation of electron microscopy (EM) images into cellular membranes. Existing pipelines typically generate a Membrane Detection Probability Map (MDPM) using machine‑learning classifiers (e.g., Random Forests, U‑Net) that assign each pixel a probability of belonging to a membrane. While MDPMs provide a useful soft‑segmentation cue, they often contain local misclassifications, noisy boundaries, and fragmented thin membranes, which limit downstream segmentation quality.
To overcome these shortcomings, the authors propose an iterative refinement framework built around a convolutional neural network (CNN) trained specifically to “recover” the label of the central pixel of a patch sampled from an MDPM. The training data consist of pairs (MDPM‑patch, true central label) where the patch is extracted from an MDPM generated by any upstream method, and the true label comes from expert‑annotated ground truth. By formulating the problem as a binary classification task (membrane vs. non‑membrane), the network learns a mapping from the local probability context to a more accurate central probability.
Network architecture and training
The CNN receives a single‑channel MDPM patch (typical sizes 65×65 or 101×101) and processes it through 4–5 convolutional layers (3×3 kernels), each followed by batch normalization and ReLU activation. A final 1×1 convolution and sigmoid activation produce a scalar probability for the central pixel. The loss function is binary cross‑entropy, optionally weighted to counter class imbalance. Training employs the Adam optimizer (learning rate ≈1e‑4), data augmentation (rotations, flips, Gaussian noise), and early stopping based on validation loss.
Iterative application
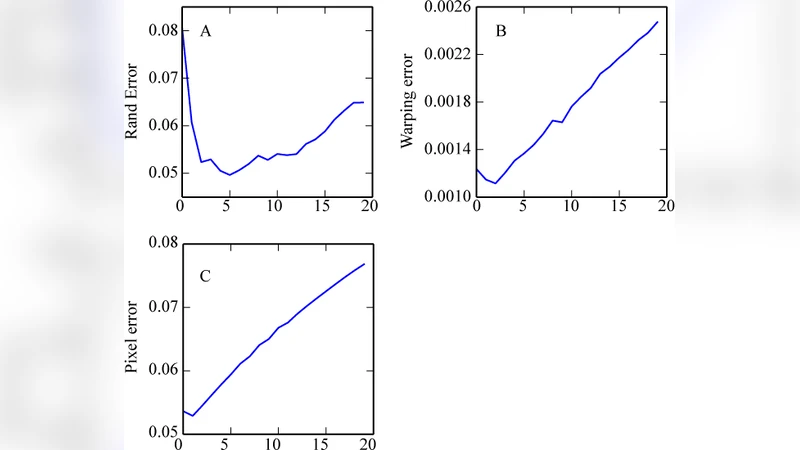
Once trained, the network is applied to the entire MDPM using a sliding‑window approach. For each pixel, the network predicts an updated probability, which replaces the original value in a new MDPM. This updated map is then fed back into the same network for another iteration. The process is repeated 3–5 times. Each iteration smooths out isolated errors, reinforces consistent membrane contours, and especially improves detection of very thin or intersecting membranes. The authors note that performance plateaus after about five passes; additional iterations yield diminishing returns while increasing computational cost.
Experimental validation
The method is evaluated on the publicly available ISBI 2012 EM dataset, a standard benchmark for membrane segmentation. Baselines include the raw MDPM (single pass), Random Forest, U‑Net, and a conventional watershed post‑processing pipeline. Evaluation metrics are the adapted Rand index (F‑Score), Variation of Information (VOI), and mean cross‑entropy (ACE). After three to five iterative passes, the proposed approach achieves an F‑Score increase from ~0.86 (baseline) to ~0.92, VOI reduction from 0.31 to 0.22, and ACE reduction from 0.12 to 0.07. Ablation studies demonstrate that a 65×65 patch size and a four‑layer network provide the best trade‑off between accuracy and runtime.
Discussion of strengths and limitations
The key contribution is the reconceptualization of MDPM as a mutable intermediate representation that can be refined through a learned feedback loop, rather than a static output awaiting a separate post‑processing step. The iterative CNN operates locally, yet its repeated application yields global consistency comparable to Markov Random Field (MRF) inference but with far lower computational overhead because all parameters are embedded in the CNN. However, the current implementation processes 2‑D slices independently, ignoring inter‑slice continuity in volumetric EM stacks. Extending the framework to 3‑D convolutions or integrating recurrent units could further improve performance on full volumes. Moreover, each iteration adds memory and compute demands; practical deployment on terabyte‑scale datasets will require careful GPU memory management and possibly model compression.
Conclusion
The authors present a simple yet powerful iterative refinement algorithm that substantially improves membrane segmentation from EM images. By training a CNN to predict the central pixel label from its surrounding probability context and repeatedly applying this predictor, the method reduces noise, repairs broken membranes, and yields higher‑quality segmentations than state‑of‑the‑art single‑pass approaches. This work demonstrates that learned, iterative post‑processing can be an effective complement to existing deep‑learning segmentation pipelines, opening avenues for more accurate and scalable connectomics reconstructions.