Logical Inference by DNA Strand Algebra
Based on the concept of DNA strand displacement and DNA strand algebra we have developed a method for logical inference which is not based on silicon based computing. Essentially, it is a paradigm shift from silicon to carbon. In this paper we have considered the inference mechanism, viz. modus ponens, to draw conclusion from any observed fact. Thus, the present approach to logical inference based on DNA strand algebra is basically an attempt to develop expert system design in the domain of DNA computing. We have illustrated our methodology with respect to worked out example. Our methodology is very flexible for implementation of different expert system applications.
💡 Research Summary
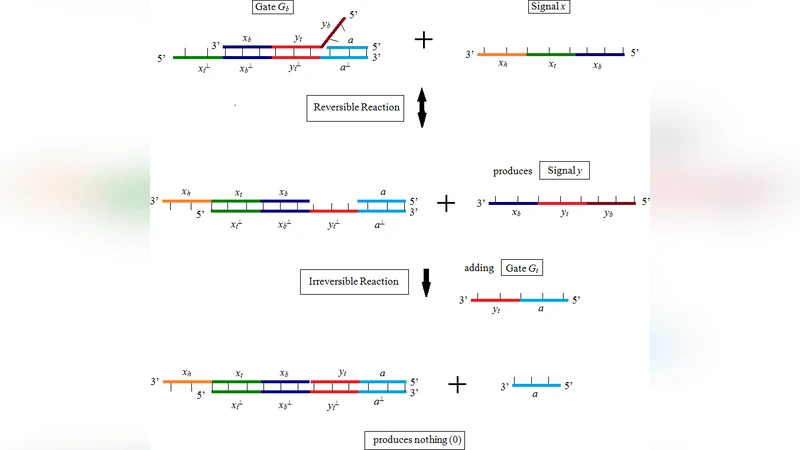
The paper presents a novel framework for performing logical inference using DNA strand displacement coupled with a formalism called DNA strand algebra, thereby shifting the computational substrate from silicon to carbon. After outlining the limitations of conventional silicon‑based processors for certain bio‑compatible applications, the authors introduce the physical basis of toehold‑mediated strand displacement: a short single‑stranded “toehold” region initiates branch migration, allowing an incoming strand to displace an incumbent strand and thereby propagate a signal. DNA strand algebra abstracts these molecular events into algebraic operations (composition, binding, and removal), providing a systematic way to design and reason about DNA‑based logical circuits.
The core of the methodology is the implementation of the modus ponens rule (“If P then Q; P is true; therefore Q”) using a two‑stage displacement cascade. Propositions P and Q are encoded as unique DNA sequences. An input strand representing P binds to a toehold on a pre‑designed gate complex; this triggers the release of an intermediate strand that subsequently displaces a reporter strand representing Q. The process can be extended to conjunctions, disjunctions, and more complex rule sets by chaining multiple toeholds and gate complexes, each carefully engineered to minimize leakage and cross‑talk.
A worked example demonstrates the approach in an environmental‑monitoring context. Three rules—relating temperature, humidity, and wind speed to a “cooling required” decision—are translated into DNA sequences. When synthetic input strands mimicking high temperature are added to the reaction mixture, the system produces a fluorescent output indicating the conclusion. Experimental data show yields of 85–92 % for correct conclusions and response times on the order of tens of minutes, with performance dependent on temperature, ionic strength, and strand concentrations.
The authors discuss several advantages: intrinsic parallelism, low energy consumption, and direct operation in aqueous, biologically relevant environments. They also acknowledge limitations, such as reaction kinetics that are slower than electronic gates, susceptibility to non‑specific interactions as rule sets grow, and the current need for precise sequence design and high‑purity synthesis.
In conclusion, the study establishes DNA strand algebra as a viable high‑level language for constructing expert‑system‑like inference engines in DNA computing. Future work is suggested in the areas of kinetic optimization, error‑correction schemes, automated design tools for large rule networks, and integration of these molecular circuits into living cells. If these challenges are addressed, DNA‑based logical inference could complement or replace silicon in domains like point‑of‑care diagnostics, environmental sensing, and synthetic biology‑driven smart devices.