Geometric friction directs cell migration
In the absence of environmental cues, a migrating cell performs an isotropic random motion. Recently, the breaking of this isotropy has been observed when cells move in the presence of asymmetric adhesive patterns. However, up to now the mechanisms at work to direct cell migration in such environments remain unknown. Here, we show that a non-adhesive surface with asymmetric micro-geometry consisting of dense arrays of tilted micro-pillars can direct cell motion. Our analysis reveals that most features of cell trajectories, including the bias, can be reproduced by a simple model of active Brownian particle in a ratchet potential, which we suggest originates from a generic elastic interaction of the cell body with the environment. The observed guiding effect, independent of adhesion, is therefore robust and could be used to direct cell migration both in vitro and in vivo.
💡 Research Summary
In this study the authors demonstrate that cells can acquire a persistent directional bias solely from the geometry of a non‑adhesive micro‑structured surface, without any chemical or biochemical cues. They fabricated dense arrays of tilted micro‑pillars in poly‑dimethyl‑siloxane (PDMS) using soft lithography, coating the pillars with polyethylene glycol (PEG) to eliminate adhesion. The pillars were 2 µm in diameter, with independently varied length (h), spacing (d) and tilt angle (α). Normal human dermal fibroblasts (NHDFs) were seeded on a fibronectin‑coated flat PDMS surface and then confined between this surface and the pillar‑bearing pad, so that the cells adhered only to the bottom surface while their nuclei contacted the non‑adhesive pillars from above. Time‑lapse imaging over 33 h allowed automatic tracking of nuclear positions, providing high‑resolution trajectories for hundreds of cells.
Two robust features emerged from the trajectories. First, cells tended to follow the two orthogonal lattice axes of the pillar array (orientation bias). Second, and more strikingly, cells migrated preferentially in the direction of the pillar tilt (forward, Fwd) rather than opposite (backward, Bwd). This direction bias was observed for all tested geometries and persisted when the bottom substrate was a soft poly‑acrylamide gel (15 kPa), indicating that the effect is not dependent on substrate stiffness. The bias increased with pillar density and tilt angle, reaching up to 78 % of cells moving forward after four hours for d = 5 µm, h = 5 µm and α = 45°. In contrast, straight (non‑tilted) pillars produced only orientation bias and no direction bias.
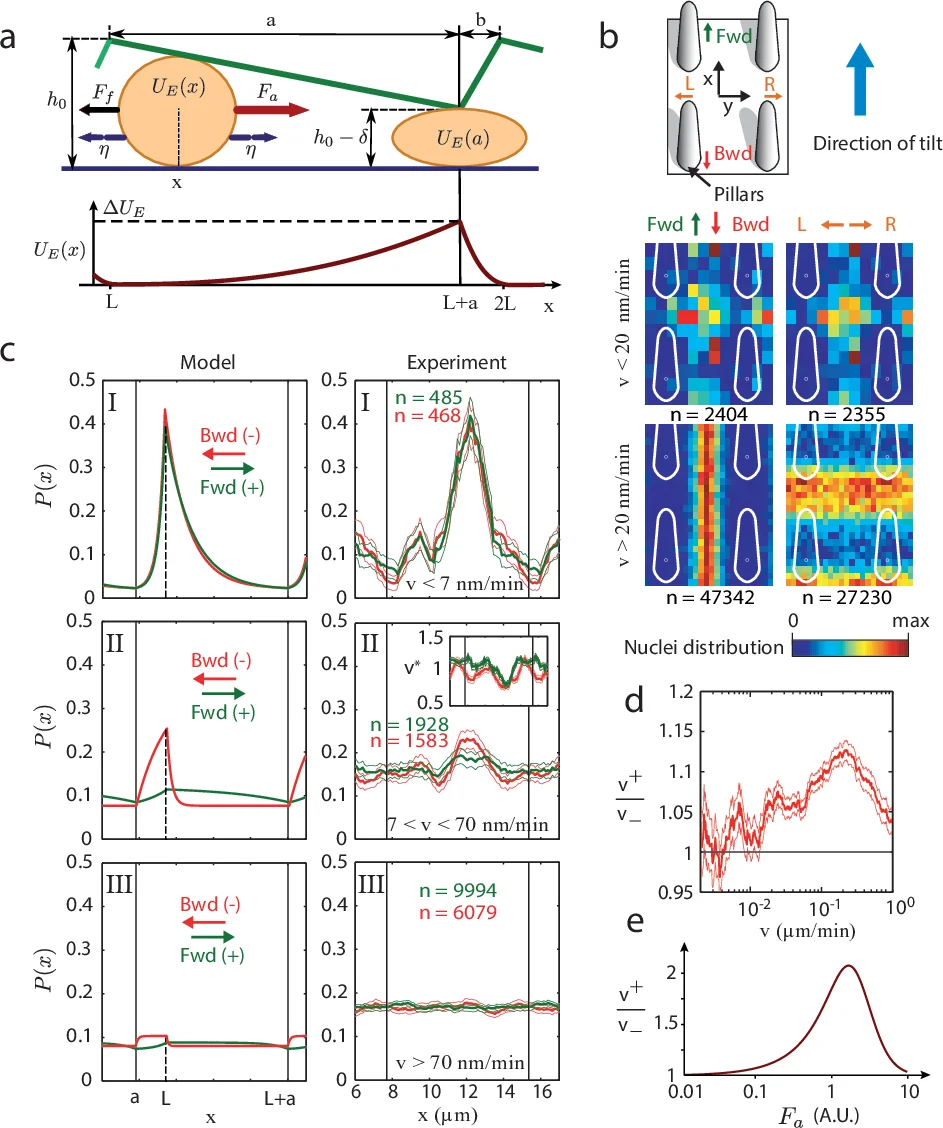
To uncover the physical origin, the authors modeled the cell as an active Brownian particle moving in a one‑dimensional ratchet potential generated by the asymmetric pillar profile. The cell nucleus was approximated as a spherical elastic body of radius R and Young’s modulus E. When the local pillar height h(x) becomes smaller than the nuclear diameter (2R), the nucleus must deform, storing elastic energy U_E(x) ∝ E R^{1/2}(2R − h)^{5/2}. This creates periodic energy barriers that are higher when the cell moves against the tilt (Bwd) than when it moves with the tilt (Fwd). The cell exerts a constant active propulsion force F_a = F + η(t), where η(t) is fast white noise, and experiences linear viscous friction λ. The overdamped Langevin equation λ dx/dt = −∂_xU_E + η leads to a tilted effective potential Ũ(x)=U_E(x)−F x.
Analysis of this model yields three dynamical regimes depending on the ratio of the maximal elastic barrier ΔU_E to the work done by the active force over one period, F L (L = a + b, the spatial period of the sawtooth). (I) ΔU_E ≫ F L: barriers dominate, motion is essentially blocked in both directions, and no bias appears. (II) ΔU_E ≈ F L: barriers are comparable to the active work; the forward direction experiences a reduced effective barrier (ΔU_E − F a) while the backward direction faces a larger one (ΔU_E − F b). This asymmetry produces a measurable velocity ratio v⁺/v⁻ > 1 and a net drift (geometric friction). (III) ΔU_E ≪ F L: barriers are negligible, the particle behaves isotropically and the bias disappears.
The authors validated the model by constructing probability density maps P(x,y) of nuclear positions relative to the surrounding pillars, sorted by instantaneous speed and direction. For intermediate speeds (7–70 nm min⁻¹, corresponding to regime II) the density peaks at the minimal pillar height, but the peak is markedly higher for forward motion, and the local speed drops, indicating that nuclei are temporarily trapped at the bottlenecks. The experimentally measured forward‑to‑backward speed ratio v⁺/v⁻ as a function of median speed reproduces the non‑monotonic curve predicted by the theory. Similar results were obtained with micro‑prisms, confirming that the effect is generic to asymmetric, non‑adhesive micro‑geometries.
In summary, the paper provides compelling experimental evidence and a quantitative theoretical framework showing that purely geometric, non‑adhesive micro‑structures generate an asymmetric energy landscape that biases active cellular motion. This “geometric friction” operates without any specific molecular adhesion, making it a robust and tunable tool for directing cell migration in vitro and potentially in vivo. The findings open new avenues for engineering tissue scaffolds, guiding immune cell trafficking, or controlling metastatic cell invasion by designing appropriate micro‑geometries rather than relying on biochemical gradients.
Comments & Academic Discussion
Loading comments...
Leave a Comment