Spectral Clustering with Epidemic Diffusion
Spectral clustering is widely used to partition graphs into distinct modules or communities. Existing methods for spectral clustering use the eigenvalues and eigenvectors of the graph Laplacian, an operator that is closely associated with random walks on graphs. We propose a new spectral partitioning method that exploits the properties of epidemic diffusion. An epidemic is a dynamic process that, unlike the random walk, simultaneously transitions to all the neighbors of a given node. We show that the replicator, an operator describing epidemic diffusion, is equivalent to the symmetric normalized Laplacian of a reweighted graph with edges reweighted by the eigenvector centralities of their incident nodes. Thus, more weight is given to edges connecting more central nodes. We describe a method that partitions the nodes based on the componentwise ratio of the replicator’s second eigenvector to the first, and compare its performance to traditional spectral clustering techniques on synthetic graphs with known community structure. We demonstrate that the replicator gives preference to dense, clique-like structures, enabling it to more effectively discover communities that may be obscured by dense intercommunity linking.
💡 Research Summary
The paper introduces a novel spectral clustering framework that replaces the traditional random‑walk‑based diffusion with epidemic diffusion, modeled by the replicator operator. In a random walk, a walker moves to a single neighbor at each step, and the graph Laplacian (or its normalized version) captures this stochastic process. By contrast, an epidemic simultaneously “infects” all neighbors of a node, leading to a non‑conservative replication of the diffusing quantity. Lerman and Ghosh previously defined the replicator matrix R = λ_max I − A, where A is the adjacency matrix and λ_max is its largest eigenvalue; the corresponding eigenvector θ is the eigenvector centrality of the graph.
The authors’ key theoretical contribution is to show that the replicator is exactly equivalent to the symmetric normalized Laplacian of a re‑weighted graph. The re‑weighting multiplies each original edge weight A_ij by the product of the eigenvector centralities of its endpoints, yielding a new adjacency matrix \tilde A_ij = A_ij θ_i θ_j. The degree of node i in this re‑weighted graph becomes \tilde d_i = λ_max θ_i², which can be expressed compactly as \tilde D = λ_max Θ² where Θ is the diagonal matrix of θ. Substituting these into the definition of the symmetric normalized Laplacian L̃_s = I − \tilde D^{‑1/2}\tilde A\tilde D^{‑1/2} leads to L̃_s = I − (1/λ_max) A, and therefore R = λ_max L̃_s. This equivalence bridges epidemic diffusion and the well‑studied diffusion of random walks, but on a graph whose edges have been amplified according to node centrality. Consequently, edges linking highly central nodes become expensive to cut, while peripheral connections remain cheap.
Leveraging this equivalence, the authors devise an efficient spectral partitioning algorithm. They compute the first two eigenvectors of the replicator: the dominant eigenvector θ (centrality) and the second eigenvector ψ. For each node i they form the component‑wise ratio v_i = ψ_i / θ_i, sort the nodes by v_i, and examine all N − 1 possible binary cuts induced by this ordering. The quality of each cut is measured by the normalized cut on the re‑weighted graph, denoted \tilde N(S). The cut minimizing \tilde N(S) is selected as the final bipartition. This procedure mirrors classic spectral bisection (which uses the second eigenvector of the Laplacian) but replaces the simple eigenvector sign test with the ψ/θ ratio, and replaces the original graph’s cut cost with the re‑weighted normalized cut. Because the ratio ψ/θ coincides with the second eigenvector of the random‑walk Laplacian L_rw, the algorithm can be implemented with the same computational tools used for standard spectral clustering, preserving the O(N log N) efficiency of eigen‑decomposition plus sorting.
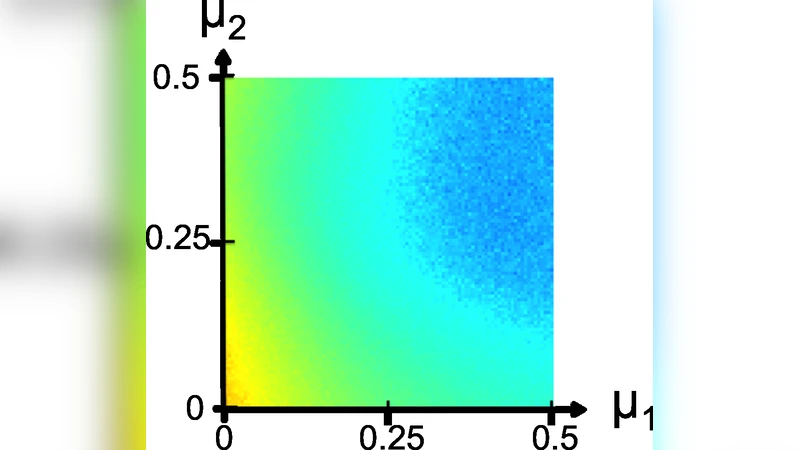
Experimental evaluation focuses on synthetic hierarchical graphs generated by the Lancichinetti–Fortunato benchmark. Nodes are organized into macro‑communities, each containing micro‑communities; two mixing parameters µ₁ (inter‑macro edge fraction) and µ₂ (inter‑micro edge fraction) control the density of inter‑community links. As µ₁ and µ₂ increase, traditional spectral methods based on L (unnormalized Laplacian) or L_s (symmetric normalized Laplacian) experience sharp declines in Normalized Mutual Information (NMI) and clustering accuracy because dense inter‑community edges blur community boundaries. In contrast, the replicator‑based method retains high NMI and accuracy even for large µ₁, demonstrating robustness to noisy inter‑community connections.
A concrete illustrative example further clarifies the mechanism. A small graph consists of a dense clique linked to a sparse cluster through a high‑degree hub node. Standard normalized cut on the original graph prefers a cut that isolates the hub with the sparse side, ignoring the dense clique structure. After re‑weighting edges by centrality, the hub’s connections to the clique become heavily weighted, making the cut that separates the hub from the clique far more expensive. Consequently, the optimal cut on the re‑weighted graph groups the hub with the clique, preserving the dense subgraph. Numerical values of ratio cut and normalized cut before and after re‑weighting confirm this behavior.
In summary, the paper establishes that epidemic diffusion, captured by the replicator, is mathematically identical to diffusion on a centrality‑re‑weighted graph. This insight enables a spectral clustering algorithm that naturally emphasizes dense, central structures (cliques) and is less sensitive to noisy inter‑community edges. The method is computationally comparable to existing spectral techniques, yet yields superior community recovery on graphs where high‑centrality hub nodes obscure true modular organization. The authors suggest future work on applying the approach to real‑world large‑scale social networks and extending the framework to dynamic or temporal graphs.
Comments & Academic Discussion
Loading comments...
Leave a Comment