Correlation dimension of complex networks
We propose a new measure to characterize the dimension of complex networks based on the ergodic theory of dynamical systems. This measure is derived from the correlation sum of a trajectory generated by a random walker navigating the network, and extends the classical Grassberger-Procaccia algorithm to the context of complex networks. The method is validated with reliable results for both synthetic networks and real-world networks such as the world air-transportation network or urban networks, and provides a computationally fast way for estimating the dimensionality of networks which only relies on the local information provided by the walkers.
💡 Research Summary
The paper introduces a novel framework for quantifying the dimensionality of complex networks by extending the classical Grassberger‑Procaccia correlation‑dimension method to discrete graph structures. The authors observe that traditional notions of dimension are rooted in continuous Euclidean spaces and therefore do not translate directly to networks, which are inherently discrete and often lack a regular geometry. To bridge this gap, they draw on ergodic theory and dynamical‑systems concepts, constructing a trajectory on the network through a simple random walk. Because a random walk on a connected graph is ergodic, a sufficiently long walk samples the entire network uniformly, regardless of the starting node. This trajectory provides a time‑ordered sequence of visited vertices that can be treated as points in an abstract state space.
For the generated sequence {x(t)} the authors compute the correlation sum
C(r) = (2 / N(N‑1)) Σ_{i<j} Θ(r – d(x_i, x_j)),
where Θ is the Heaviside step function and d(·,·) denotes a distance metric defined on the graph. In practice the metric is taken as the shortest‑path length (or a suitably embedded metric) between two vertices. By plotting log C(r) versus log r and identifying a scaling region where C(r) ∝ r^{D_2}, the slope D_2 provides an estimate of the network’s correlation dimension, denoted D_2^net. This procedure mirrors the original Grassberger‑Procaccia algorithm but replaces Euclidean distances with graph‑theoretic distances, thereby preserving the non‑Euclidean nature of the underlying structure.
The authors first validate the approach on synthetic networks. Random Erdős‑Rényi graphs yield very large or diverging D_2 values, reflecting their lack of geometric constraints. Regular lattices in two and three dimensions recover D_2≈2 and D_2≈3 respectively, confirming that the method correctly identifies known embedding dimensions. Scale‑free Barabási‑Albert networks produce intermediate dimensions, illustrating how hub‑centric connectivity reduces the effective dimensionality. Small‑world Watts‑Strogatz graphs exhibit a smooth transition of D_2 as the rewiring probability varies, demonstrating sensitivity to the balance between local clustering and long‑range shortcuts.
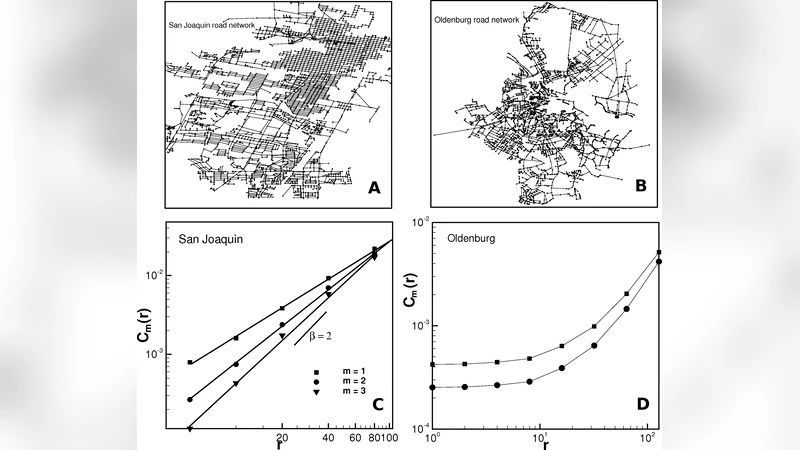
Real‑world applications include the worldwide air‑transportation network and several urban transportation graphs (road and public‑transit networks). The air‑transport network shows D_2≈2.5, suggesting that while the system is embedded on a roughly planar Earth surface, the presence of hub airports and long‑haul flights adds a fractional dimensional component. Urban networks typically yield D_2 values between 1.8 and 2.2, consistent with a primarily two‑dimensional layout but slightly elevated by multilayer infrastructure such as overpasses and subways.
A systematic analysis of methodological parameters is provided. The length of the random‑walk trajectory N must be large enough (10^5–10^6 steps in the experiments) to achieve statistical stability; shorter walks lead to noisy C(r) curves and biased dimension estimates, especially at small radii. The choice of the scaling interval for r is crucial: at radii smaller than the average shortest‑path length, the correlation sum reflects local clustering and triangle density, whereas at larger radii it captures global, possibly fractal‑like scaling. The authors demonstrate that many networks exhibit distinct scaling regimes, implying that a single scalar dimension may be insufficient to describe their full geometric complexity.
From a computational standpoint, the algorithm requires only one random‑walk trajectory and uses efficient data structures (KD‑trees or ball‑trees) to compute pairwise distances, achieving O(N log N) time and O(N) memory. This is a substantial improvement over methods that need the full all‑pairs shortest‑path matrix (O(N^2) memory), making the approach scalable to graphs with millions of nodes and suitable for online or streaming contexts where only local information is available.
In conclusion, the paper delivers a fast, locally‑driven, and theoretically grounded technique for estimating the correlation dimension of complex networks. By leveraging the ergodic nature of random walks, it transforms a purely topological object into a dynamical dataset amenable to classic nonlinear‑time‑series analysis. The extensive validation on both synthetic and empirical networks confirms the method’s accuracy and robustness. Future directions suggested include applying alternative dynamical processes (e.g., diffusion, routing) to generate trajectories, extending the framework to temporal networks to track dimensional evolution, and integrating the dimension estimate with other network descriptors for richer characterizations of complexity.