Circuit architecture explains functional similarity of bacterial heat shock responses
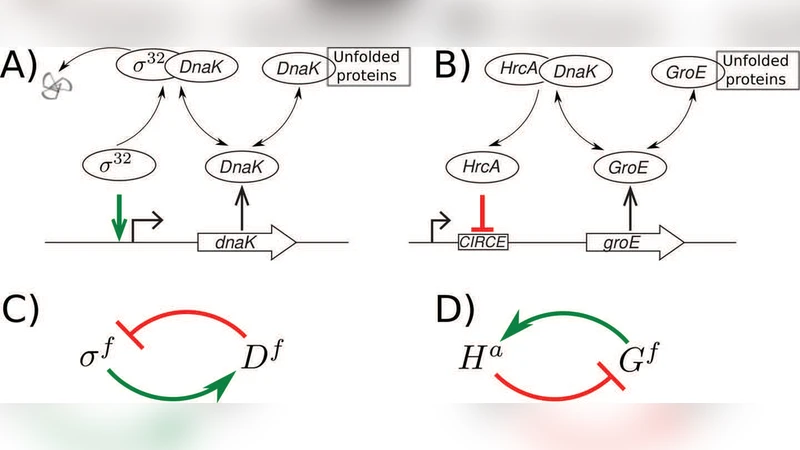
Heat shock response is a stress response to temperature changes and a consecutive increase in amounts of unfolded proteins. To restore homeostasis, cells upregulate chaperones facilitating protein folding by means of transcription factors (TF). We here investigate two heat shock systems: one characteristic to gram negative bacteria, mediated by transcriptional activator sigma32 in E. coli, and another characteristic to gram positive bacteria, mediated by transcriptional repressor HrcA in L. lactis. We construct simple mathematical model of the two systems focusing on the negative feedbacks, where free chaperons suppress sigma32 activation in the former, while they activate HrcA repression in the latter. We demonstrate that both systems, in spite of the difference at the TF regulation level, are capable of showing very similar heat shock dynamics. We find that differences in regulation impose distinct constrains on chaperone-TF binding affinities: the binding constant of free sigma32 to chaperon DnaK, known to be in 100 nM range, set the lower limit of amount of free chaperon that the system can sense the change at the heat shock, while the binding affinity of HrcA to chaperon GroE set the upper limit and have to be rather large extending into the micromolar range.
💡 Research Summary
The paper presents a comparative systems‑biology analysis of two canonical bacterial heat‑shock response (HSR) circuits: the σ32‑driven system of the Gram‑negative bacterium Escherichia coli and the HrcA‑driven system of the Gram‑positive Lactococcus lactis. Although the transcriptional regulators operate through opposite mechanisms—σ32 is an activator whose activity is inhibited by the chaperone DnaK, whereas HrcA is a repressor whose activity is relieved by the chaperone GroE—the authors show that both networks generate remarkably similar dynamic responses to temperature up‑shifts.
To expose the core principles, the authors construct minimal ordinary‑differential‑equation (ODE) models that retain only the essential negative‑feedback loops. In the σ32 model, free DnaK binds σ32, forming an inactive complex; heat shock increases the pool of unfolded proteins, sequestering DnaK and thereby freeing σ32 to up‑regulate the expression of DnaK, GroEL/GroES, and other chaperones. In the HrcA model, free GroE binds HrcA, preventing HrcA from repressing the groE operon; heat shock again raises unfolded protein levels, which consume free GroE, releasing HrcA from inhibition and allowing transcription of GroE and other chaperones. Both models include synthesis and degradation terms for the transcription factor, the chaperone, and the unfolded‑protein load, as well as binding equilibria characterized by dissociation constants (Kd).
Parameter analysis reveals that the two circuits impose opposite constraints on the TF‑chaperone binding affinity. For the σ32‑DnaK interaction, experimental measurements place Kd in the low‑nanomolar range (~100 nM). The model demonstrates that such a high affinity sets a lower detection limit for free DnaK: the system can sense only a modest reduction in free DnaK before σ32 becomes fully active, ensuring a rapid and sensitive response without excessive noise. Conversely, the HrcA‑GroE interaction must be considerably weaker. Simulations indicate that Kd values in the micromolar range (≈1–10 µM) are required for the circuit to function properly. A tighter binding would keep HrcA permanently repressed, blunting the heat‑shock response, whereas a very weak interaction would make the system overly permissive, leading to constitutive chaperone expression. Thus, the HrcA circuit imposes an upper limit on the affinity that must be satisfied for proper regulation.
Dynamic simulations of both models under a step increase in temperature produce the classic “fast rise – slow decay” profile of chaperone concentrations: a rapid surge in DnaK or GroE followed by a gradual return to baseline as unfolded proteins are refolded. Despite the opposite regulatory logic, the time courses, peak amplitudes, and relaxation times are nearly indistinguishable when the respective affinity constraints are respected. This convergence underscores that the architecture of a negative‑feedback loop, rather than the polarity of transcriptional control, dictates the overall system dynamics.
The authors discuss the evolutionary implications of this finding. The convergence of functional dynamics despite divergent molecular mechanisms suggests that bacterial lineages have arrived at a common design principle: a chaperone‑sensing feedback that buffers proteostasis against thermal stress. Moreover, the work offers practical guidance for synthetic biology. When engineering heat‑shock circuits in heterologous hosts, one can achieve desired response characteristics by tuning the TF‑chaperone Kd to the appropriate regime (low‑nanomolar for activator‑based designs, micromolar for repressor‑based designs). This provides a quantitative target for protein‑engineering efforts aimed at constructing temperature‑responsive gene‑expression modules, stress‑tolerant production strains, or programmable biosensors.
In summary, the study demonstrates that the σ32‑DnaK and HrcA‑GroE circuits, though mechanistically opposite, are functionally equivalent because they share a core negative‑feedback topology. The key distinction lies in the required binding affinities: σ32 must bind DnaK tightly to set a low detection threshold, whereas HrcA must bind GroE weakly to keep the detection threshold high. This insight deepens our understanding of bacterial stress‑response architecture and equips researchers with concrete design parameters for future applications in systems and synthetic biology.
Comments & Academic Discussion
Loading comments...
Leave a Comment