Geometric Tomography With Topological Guarantees
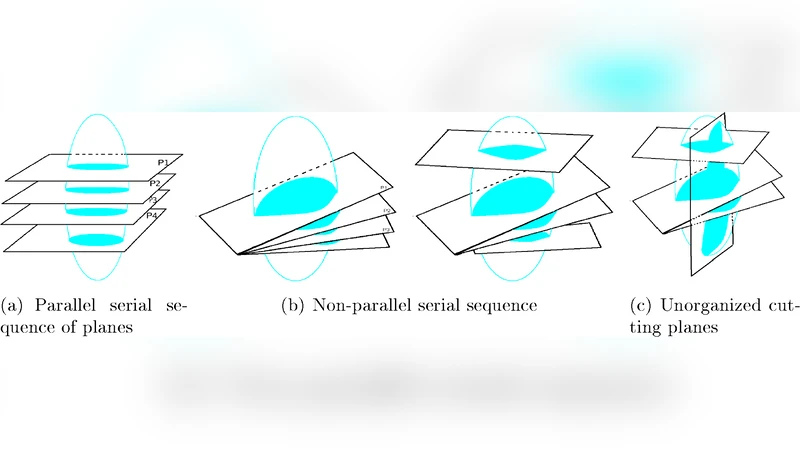
We consider the problem of reconstructing a compact 3-manifold (with boundary) embedded in $\mathbb{R}^3$ from its cross-sections $\mathcal S$ with a given set of cutting planes $\mathcal P$ having arbitrary orientations. Using the obvious fact that a point $x \in \mathcal P$ belongs to the original object if and only if it belongs to $\mathcal S$, we follow a very natural reconstruction strategy: we say that a point $x \in \mathbb{R}^3$ belongs to the reconstructed object if (at least one of) its nearest point(s) in $\mathcal P$ belongs to $\mathcal S$. This coincides with the algorithm presented by Liu et al. in \cite{LB+08}. In the present paper, we prove that under appropriate sampling conditions, the output of this algorithm preserves the homotopy type of the original object. Using the homotopy equivalence, we also show that the reconstructed object is homeomorphic (and isotopic) to the original object. This is the first time that 3-dimensional shape reconstruction from cross-sections comes with theoretical guarantees.
💡 Research Summary
The paper addresses the classic problem of reconstructing a compact three‑dimensional manifold with boundary from a collection of planar cross‑sections. Given a set 𝒫 of cutting planes with arbitrary orientations and the corresponding intersection curves 𝒮, the authors adopt the most natural reconstruction rule: a point x∈ℝ³ belongs to the reconstructed object R if the nearest point of x on any plane p∈𝒫 lies on the measured cross‑section Sₚ. This rule coincides with the algorithm introduced by Liu et al. (2008), but the present work provides a rigorous topological analysis that was previously missing.
The core contribution is a set of sampling conditions under which the output R is guaranteed to have the same homotopy type as the original manifold M, and, moreover, to be homeomorphic and isotopic to M. The conditions are expressed in terms of the reach ρ of M (the minimum distance to its medial axis) and an ε‑sampling density: every point of M must be within ε·ρ of at least one cutting plane, with ε<½. Additionally, the minimal distance δ between any two planes must satisfy δ≤ε·ρ. Under these constraints the authors construct continuous maps f:R→M and g:M→R that are mutual homotopy inverses. The proof relies on the Nerve theorem: the family of ε‑balls around the planes forms an open cover of M whose nerve is combinatorially equivalent to the intersection pattern of the cross‑sections, and this nerve is also the nerve of the reconstructed volume. Consequently, the nerve complexes of R and M are isomorphic, establishing a homotopy equivalence.
Beyond homotopy, the authors show that the constructed maps can be refined to yield a homeomorphism, and that a smooth isotopy exists between R and M, meaning one can be continuously deformed into the other without tearing or self‑intersection. The paper also discusses algorithmic aspects: nearest‑plane queries can be answered efficiently using spatial data structures (k‑d trees, distance transforms), leading to an overall reconstruction time that scales linearly with the volume resolution and logarithmically with the number of planes.
Experimental validation is performed on synthetic models and medical CT data. By varying the plane density and the ε parameter, the authors demonstrate that when the theoretical sampling bounds are respected, the reconstructed volumes match the original topology exactly, while violations of the bounds produce spurious holes or extra components.
In summary, this work provides the first theoretical guarantees for three‑dimensional shape reconstruction from arbitrary planar cross‑sections. It bridges a gap between practical reconstruction pipelines and rigorous topological correctness, opening the door for reliable applications in medical imaging, cultural heritage preservation, and industrial inspection where topological fidelity is critical. Future work is suggested to relax the sampling assumptions, handle partial or noisy sections, and extend the framework to point‑cloud based scanning modalities.
Comments & Academic Discussion
Loading comments...
Leave a Comment