Optimal reconstruction of the folding landscape using differential energy surface analysis
In experiments and in simulations, the free energy of a state of a system can be determined from the probability that the state is occupied. However, it is often necessary to impose a biasing potentia
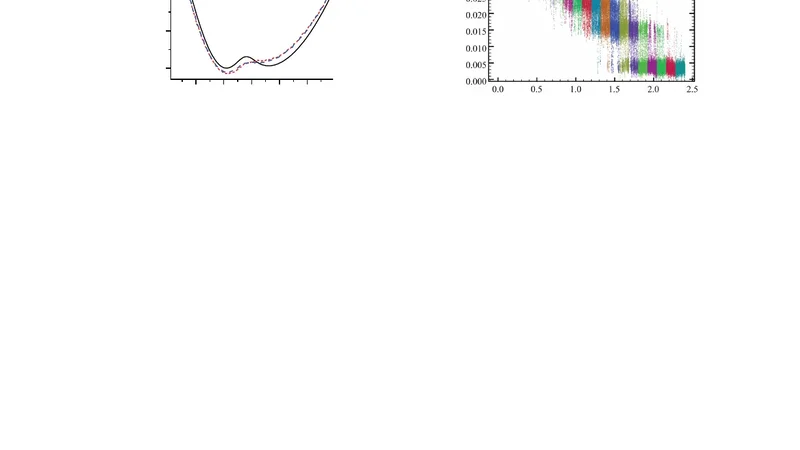
In experiments and in simulations, the free energy of a state of a system can be determined from the probability that the state is occupied. However, it is often necessary to impose a biasing potential on the system so that high energy states are sampled with sufficient frequency. The unbiased energy is typically obtained from the data using the weighted histogram analysis method (WHAM). Here we present differential energy surface analysis (DESA), in which the gradient of the energy surface, dE/dx, is extracted from data taken with a series of harmonic biasing potentials. It is shown that DESA produces a maximum likelihood estimate of the folding landscape gradient. DESA is demonstrated by analyzing data from a simulated system as well as data from a single-molecule unfolding experiment in which the end-to-end distance of a DNA hairpin is measured. It is shown that the energy surface obtained from DESA is indistinguishable from the energy surface obtained when WHAM is applied to the same data. Two criteria are defined which indicate whether the DESA results are self-consistent. It is found that these criteria can detect a situation where the energy is not a single-valued function of the measured reaction coordinate. The criteria were found to be satisfied for the experimental data analyzed, confirming that end-to-end distance is a good reaction coordinate for the experimental system. The combination of DESA and the optical trap assay in which a structure is disrupted under harmonic constraint facilitates an extremely accurate measurement of the folding energy surface.
💡 Research Summary
The paper introduces Differential Energy Surface Analysis (DESA), a method for reconstructing the free‑energy landscape of a molecular system from data collected under a series of harmonic biasing potentials. Traditional approaches such as the Weighted Histogram Analysis Method (WHAM) combine biased probability distributions to obtain the unbiased free energy E(x). WHAM works well when the data are abundant and the biases are comparable, but it can become cumbersome when bias strengths differ widely or when high‑energy states are poorly sampled.
DESA takes a different route: instead of estimating E(x) directly, it extracts the gradient dE/dx from each biased experiment. For a harmonic bias V_i(x)=½k_i(x−x_i^0)^2 the biased probability obeys p_i(x)∝exp
📜 Original Paper Content
🚀 Synchronizing high-quality layout from 1TB storage...