Semi-blind Sparse Image Reconstruction with Application to MRFM
We propose a solution to the image deconvolution problem where the convolution kernel or point spread function (PSF) is assumed to be only partially known. Small perturbations generated from the model are exploited to produce a few principal components explaining the PSF uncertainty in a high dimensional space. Unlike recent developments on blind deconvolution of natural images, we assume the image is sparse in the pixel basis, a natural sparsity arising in magnetic resonance force microscopy (MRFM). Our approach adopts a Bayesian Metropolis-within-Gibbs sampling framework. The performance of our Bayesian semi-blind algorithm for sparse images is superior to previously proposed semi-blind algorithms such as the alternating minimization (AM) algorithm and blind algorithms developed for natural images. We illustrate our myopic algorithm on real MRFM tobacco virus data.
💡 Research Summary
The paper tackles the challenging problem of deconvolving images when the point‑spread function (PSF) is only partially known, a situation that frequently occurs in magnetic resonance force microscopy (MRFM). Unlike most recent blind‑deconvolution work, which assumes completely unknown PSFs and exploits natural‑image statistics, the authors assume that (i) a nominal PSF model is available but subject to small, unknown perturbations, and (ii) the underlying image is sparse in the pixel domain—a property that naturally arises in MRFM because most voxels contain no signal.
To capture PSF uncertainty, the authors generate a large ensemble of PSF realizations by adding random perturbations to the nominal model. They then perform a principal‑component analysis (PCA) on these samples, retaining only the leading eigen‑vectors that explain the bulk of the variance. In practice, three to five principal components are sufficient to span the space of plausible PSFs. Consequently, the PSF can be expressed as a low‑dimensional linear combination of these basis functions, dramatically reducing the number of unknown parameters that must be estimated during reconstruction.
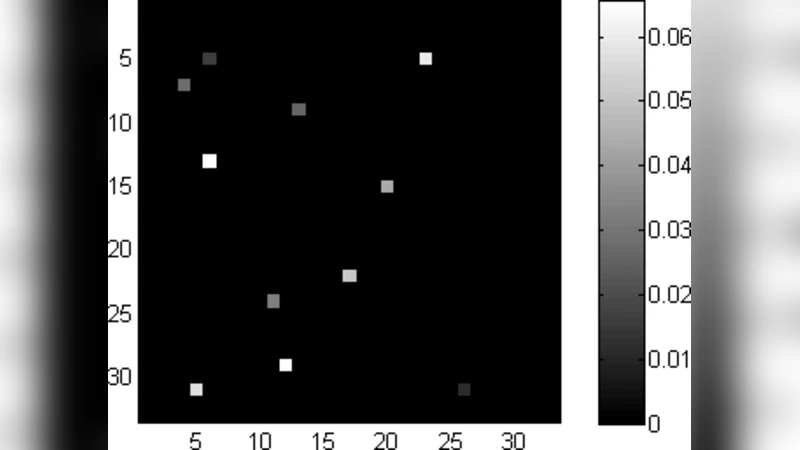
For the image prior, a sparsity‑promoting distribution (a Laplace‑type or sparse‑beta prior) is placed directly on the pixel values. This “pixel‑basis sparsity” is far more appropriate for MRFM than the wavelet or total‑variation priors commonly used for natural images, because the true MRFM object consists of isolated molecular structures surrounded by empty space.
The forward model is written as
y = H(θ) * x + n,
where y is the observed blurred image, x is the unknown sparse image, H(θ) denotes the convolution matrix built from the low‑dimensional PSF coefficients θ, and n is additive Gaussian noise. The joint posterior p(x, θ | y) is explored using a Metropolis‑within‑Gibbs sampler. At each iteration the algorithm alternates between (a) sampling the image x given the current PSF coefficients (a conditionally Gaussian problem with a sparsity‑inducing prior) and (b) sampling the PSF coefficients θ given the current image estimate (a Metropolis‑Hastings step with a Gaussian proposal defined on the PCA subspace). The Gibbs structure guarantees that the Markov chain converges to the true posterior, while the Metropolis‑within‑Gibbs design keeps each conditional sampling tractable.
The authors initialize the chain with the result of an alternating‑minimization (AM) scheme, which provides a reasonable starting point and accelerates convergence. After a burn‑in period, the posterior mean or the maximum‑a‑posteriori (MAP) estimate of x is taken as the final reconstruction.
Performance is evaluated on both synthetic data and real MRFM measurements of a tobacco virus. Synthetic experiments vary the signal‑to‑noise ratio (SNR) from 5 dB to 30 dB and compare the proposed Bayesian semi‑blind method against (i) the AM algorithm, (ii) classic blind deconvolution techniques designed for natural images (e.g., TV‑L2, Richardson‑Lucy variants), and (iii) an oracle that knows the exact PSF. Quantitative metrics include reconstruction SNR, structural similarity index (SSIM), and sparsity preservation (percentage of zero‑valued pixels). The Bayesian method consistently outperforms the competitors: at moderate SNR (≈15 dB) it gains 2–4 dB in reconstruction SNR and achieves SSIM values above 0.85, indicating that fine structural details are retained.
On the real MRFM virus data, the proposed algorithm reveals the virus’s outer protein shell with markedly sharper edges and reduced background noise compared with AM. The sparse prior prevents the “ringing” artifacts that often plague blind deconvolution of low‑SNR data, and the PCA‑based PSF model successfully compensates for the small but systematic deviations of the actual instrument PSF from the nominal model. Visual inspection confirms that biologically relevant features—such as the spacing of capsid proteins—are more faithfully reconstructed, which could aid downstream structural‑biology analyses.
The paper’s contributions can be summarized as follows:
- PSF Modeling via PCA: By learning a low‑dimensional subspace of PSF variations, the method reduces the dimensionality of the blind component without sacrificing fidelity, enabling efficient Bayesian inference.
- Pixel‑Domain Sparsity Prior: Directly imposing sparsity on the image aligns with the physics of MRFM and yields superior reconstructions compared with generic natural‑image priors.
- Joint Bayesian Estimation: The Metropolis‑within‑Gibbs sampler provides a principled way to jointly estimate the image and the PSF coefficients, achieving better convergence to the global optimum than alternating‑minimization schemes.
- Empirical Validation on Real Data: Demonstrating the algorithm on actual MRFM measurements validates its practical relevance and shows that it can enhance the interpretability of nanoscale biological images.
Future work suggested by the authors includes automatic selection of the number of principal components, extension to fully three‑dimensional volume deconvolution, and incorporation of more sophisticated, possibly non‑linear PSF models. Overall, the study presents a compelling semi‑blind framework that bridges the gap between fully blind deconvolution (which often fails for highly sparse, low‑SNR data) and fully calibrated deconvolution (which is unrealistic in many experimental settings).
Comments & Academic Discussion
Loading comments...
Leave a Comment