Network Structure, Topology and Dynamics in Generalized Models of Synchronization
We explore the interplay of network structure, topology, and dynamic interactions between nodes using the paradigm of distributed synchronization in a network of coupled oscillators. As the network evolves to a global steady state, interconnected oscillators synchronize in stages, revealing network’s underlying community structure. Traditional models of synchronization assume that interactions between nodes are mediated by a conservative process, such as diffusion. However, social and biological processes are often non-conservative. We propose a new model of synchronization in a network of oscillators coupled via non-conservative processes. We study dynamics of synchronization of a synthetic and real-world networks and show that different synchronization models reveal different structures within the same network.
💡 Research Summary
The paper addresses a fundamental question in network science: how the topology of a network and the dynamics that run on it influence each other. Traditional synchronization studies have largely relied on conservative processes such as diffusion, which are mathematically captured by the graph Laplacian and assume that the quantity exchanged between nodes is conserved. While appropriate for many physical systems, this assumption fails to describe a wide range of social, biological, and engineered processes where information, energy, or biochemical signals can be amplified, attenuated, or otherwise transformed during transmission.
To overcome this limitation, the authors introduce a generalized synchronization framework that explicitly incorporates non‑conservative interactions. Each node i carries a phase θ_i that evolves according to
dθ_i/dt = ω_i + Σ_j A_{ij} f(θ_j−θ_i) ,
where ω_i is the intrinsic frequency, A_{ij} is a directed weight matrix (allowing asymmetry), and f is a non‑linear, non‑symmetric coupling function. By choosing f to represent amplification for positive phase differences and suppression for negative ones, the model captures processes such as opinion reinforcement, neurotransmitter release, or gene‑regulatory feedback that are not energy‑preserving. The formulation reduces to the classic Kuramoto model when A is symmetric and f is sinusoidal, thereby demonstrating that the new model subsumes the conventional case.
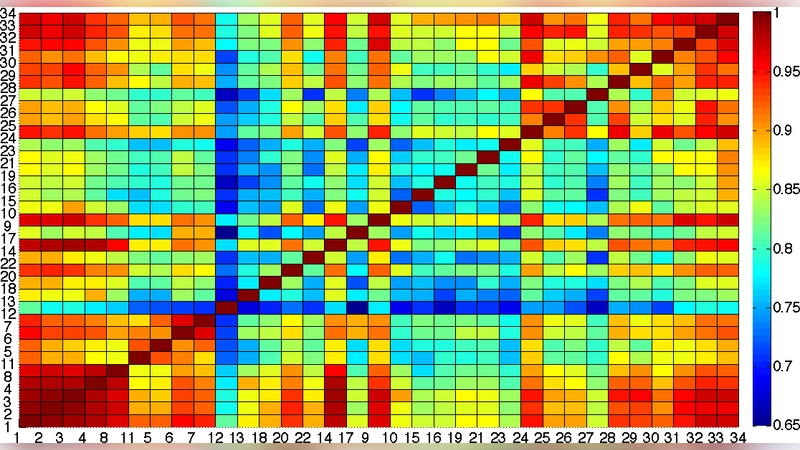
A central finding is that synchronization proceeds in discrete stages. Initially, tightly knit clusters of nodes with strong internal connections rapidly achieve local synchrony. These clusters correspond closely to the modular organization of the underlying graph. As time progresses, locally synchronized groups begin to interact, leading to larger synchronized components and eventually a global steady state. By tracking the composition of synchronized clusters over time, the authors reveal a dynamic community detection method: the temporal hierarchy of synchronization mirrors the multi‑scale community structure of the network.
The authors validate their approach on two classes of networks. The first is a synthetic hierarchical network comprising 500 nodes, average degree 6, and two levels of community organization (four large modules each containing five sub‑modules). The second class consists of real‑world data: a Twitter retweet network and a human protein‑protein interaction (PPI) network. In the synthetic case, the non‑conservative model reproduces the designed hierarchy with high fidelity; sub‑modules appear as distinct synchronization plateaus, whereas the traditional Laplacian‑based model only detects the larger modules. In the empirical networks, the new model uncovers fine‑grained groups that are invisible to standard modularity maximization. For example, in the Twitter graph, a set of users discussing a niche topic forms an early‑synchronizing cluster, while in the PPI network, proteins belonging to a specific signaling pathway synchronize faster than the rest of the network.
Parameter analysis shows that the strength of non‑conservativeness (denoted β) and the shape of the coupling function f critically affect both the speed of synchronization and the final synchrony level. Moderate β values accelerate the emergence of local synchrony, but excessively high β leads to instability: amplified signals can prevent the system from reaching a coherent global state. This non‑monotonic relationship mirrors real systems where excessive information overload or energy dissipation hampers coordinated behavior.
The contributions of the paper are threefold. First, it provides a mathematically rigorous extension of synchronization theory that accommodates non‑conservative dynamics, thereby broadening the applicability of synchronization as an analytical tool. Second, it demonstrates that the temporal evolution of synchronization can serve as a powerful, dynamics‑driven community detection technique capable of revealing multi‑scale structures that static, topology‑only methods miss. Third, it offers a systematic exploration of how model parameters influence dynamical outcomes, furnishing guidelines for applying the framework to diverse domains such as sociology, neuroscience, and systems biology.
In conclusion, the study establishes that the interplay between network topology and non‑conservative dynamics yields richer information about structural organization than conservative models alone. The proposed framework opens avenues for future work, including integration with other dynamical processes (e.g., epidemic spreading, opinion formation), real‑time monitoring of evolving networks, and the development of inference algorithms that estimate the underlying non‑conservative coupling from observed time series. By moving beyond the diffusion paradigm, the paper sets a new direction for research at the intersection of network structure, dynamics, and function.
Comments & Academic Discussion
Loading comments...
Leave a Comment