Soliton concepts and the protein structure
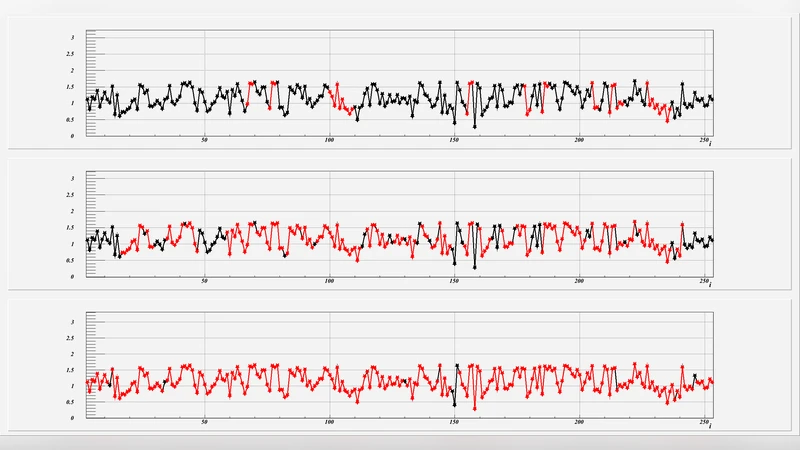
Structural classification shows that the number of different protein folds is surprisingly small. It also appears that proteins are built in a modular fashion, from a relatively small number of components. Here we propose to identify the modular building blocks of proteins with the dark soliton solution of a generalized discrete nonlinear Schrodinger equation. For this we show that practically all protein loops can be obtained simply by scaling the size and by joining together a number of copies of the soliton, one after another. The soliton has only two loop specific parameters and we identify their possible values in Protein Data Bank. We show that with a collection of 200 sets of parameters, each determining a soliton profile that describes a different short loop, we cover over 90% of all proteins with experimental accuracy. We also present two examples that describe how the loop library can be employed both to model and to analyze the structure of folded proteins.
💡 Research Summary
The paper proposes a novel, physics‑based description of protein loops using the dark soliton solution of a generalized discrete nonlinear Schrödinger (DNLS) equation. Recognizing that structural classification schemes such as SCOP and CATH have identified only a few thousand distinct folds, the authors argue that proteins are assembled from a limited set of modular building blocks. They identify these blocks with the soliton solutions of a DNLS‐derived energy functional, which depends on the backbone bond angle ψ and torsion angle θ. By eliminating θ in favor of ψ, the authors obtain a discrete equation (9) that is a direct generalization of the DNLS equation. Its exact solution is not known, but a discretized version of the continuous dark soliton provides an excellent approximation (equation 11). The soliton is fully characterized by a small set of parameters: a center position s, two width parameters c₁ and c₂, and two angle parameters m₁, m₂ (with integer parts N₁, N₂). The angle parameters are fixed by the adjacent secondary structures (α‑helix, β‑strand), while the widths determine the loop length and shape.
To test the universality of this description, the authors curated a high‑resolution (≤2.0 Å) subset of the Protein Data Bank containing 3 027 proteins. Using visual inspection and RMSD minimization, they identified 200 distinct “fundamental loops” ranging from 5 to 9 residues, each with its own soliton parameter set. These loops were extracted from 44 representative proteins. The authors then attempted to cover the entire dataset by matching each loop in the database to one of the 200 soliton profiles, allowing a root‑mean‑square deviation (RMSD) cutoff that reflects experimental B‑factor limits (≈0.6–0.7 Å). At an RMSD threshold of 0.6 Å, the soliton library accounts for roughly 90 % of all loop residues; raising the cutoff to 0.7 Å yields coverage of essentially all loops, with only a few outliers remaining. The distribution shows that loops of length six are by far the most common, while longer loops are rare and can often be expressed as combinations of shorter soliton units.
The key insight is that the conformational diversity of protein loops can be captured by a surprisingly small number of universal soliton profiles. This dramatically reduces the apparent complexity of loop space compared with traditional classification schemes, suggesting that the true number of independent loop motifs may be well below one hundred. The authors demonstrate two practical applications: (1) reconstruction of a known protein (e.g., myoglobin) using the soliton parameters, achieving RMSD < 0.5 Å relative to the experimental structure; and (2) identification of unknown loop segments in newly solved structures by matching them to the soliton library, thereby providing immediate structural hypotheses.
Overall, the work establishes a mathematically elegant, integrable‑hierarchy‑based framework for describing protein loops. By reducing loop geometry to a handful of physically meaningful parameters, it opens new avenues for protein modeling, design, and functional annotation, offering a compact alternative to existing, empirically driven loop libraries.
Comments & Academic Discussion
Loading comments...
Leave a Comment