A Novel Geographic Partitioning System for Anonymizing Health Care Data
With large volumes of detailed health care data being collected, there is a high demand for the release of this data for research purposes. Hospitals and organizations are faced with conflicting interests of releasing this data and protecting the confidentiality of the individuals to whom the data pertains. Similarly, there is a conflict in the need to release precise geographic information for certain research applications and the requirement to censor or generalize the same information for the sake of confidentiality. Ultimately the challenge is to anonymize data in order to comply with government privacy policies while reducing the loss in geographic information as much as possible. In this paper, we present a novel geographic-based system for the anonymization of health care data. This system is broken up into major components for which different approaches may be supplied. We compare such approaches in order to make recommendations on which of them to select to best match user requirements.
💡 Research Summary
The paper addresses the tension between the need to release detailed health‑care datasets for research and the legal/ethical requirement to protect patient privacy, especially when precise geographic information is involved. Traditional de‑identification removes direct identifiers but leaves quasi‑identifiers, such as demographic and location attributes, vulnerable to re‑identification through linkage attacks. Existing geographic‑based anonymization methods either apply a fixed population cut‑off (suppressing all records in regions below a threshold) or perform coarse generalization (e.g., cropping postal codes). Both approaches suffer from significant information loss: the cut‑off method can over‑suppress, while coarse generalization may unnecessarily enlarge geographic units, degrading spatial analysis.
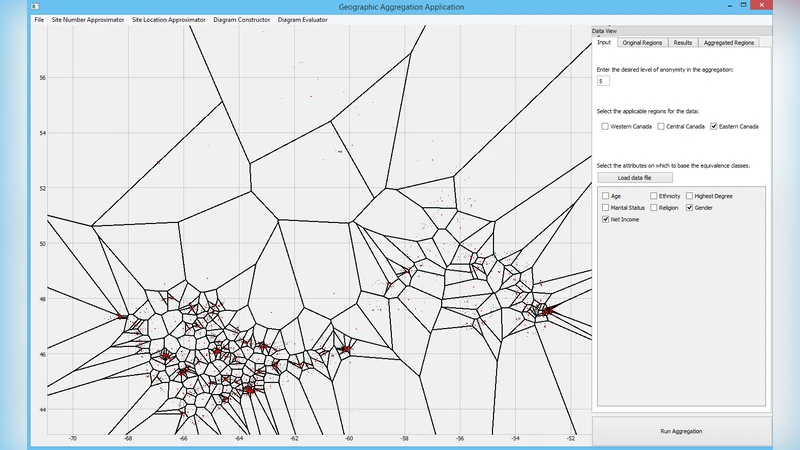
To overcome these limitations, the authors propose the Voronoi‑Based Aggregation System (VBAS), a configurable framework that uses Voronoi diagrams to partition the study area into aggregated regions that satisfy a user‑specified k‑anonymity requirement while preserving as much geographic precision as possible. VBAS is organized into four modular components:
-
Site Number Approximation – Determines how many Voronoi sites (i.e., aggregated regions) are needed. The paper evaluates several strategies, including histogram‑based population estimates, density‑based clustering, and optimization models (linear or integer programming). The goal is to choose the smallest number of sites that can still meet the k‑anonymity constraint given the distribution of quasi‑identifiers.
-
Site Location Selection – Places the chosen number of sites within the geographic space. Approaches examined include random placement, k‑means centroids, and population‑weighted centroids. The placement directly influences the shape, compactness, and population balance of the resulting Voronoi cells, which in turn affects suppression rates and utility.
-
Voronoi Construction and Aggregation – Constructs the Voronoi diagram from the selected sites and assigns each fine‑grained original region (e.g., census block) to the cell containing its centroid. If a cell’s total population is below k, the algorithm merges it with neighboring cells or applies additional suppression. This step ensures that every aggregated region meets the anonymity threshold while striving to keep cells compact and geographically coherent.
-
Rating the Aggregation – Quantifies the quality of the anonymization using several metrics:
- Suppression Rate – proportion of records removed.
- Compactness – geometric measure of how tightly the aggregated region encloses its population (e.g., area‑to‑population ratio).
- Discernibility Metric – penalizes records that become indistinguishable, reflecting loss of analytical granularity.
- Non‑Uniform Entropy – estimates information loss based on the probability of correctly guessing original attribute values from anonymized values, weighting more uniform distributions higher.
The authors implemented VBAS and evaluated it on Canadian health‑care data combined with census population statistics. Experiments compared different combinations of the four components. Key findings include:
- VBAS consistently reduced suppression rates by more than 30 % compared with a naïve cut‑off approach while only expanding average region size by roughly 20 %.
- Population‑weighted site placement produced the most compact Voronoi cells, preserving spatial resolution crucial for epidemiological studies.
- The modular design allowed the system to adapt to various k values (e.g., k = 10, 20) without manual re‑tuning of thresholds.
The paper’s contributions are threefold:
- A novel, geometry‑driven anonymization framework that dynamically determines region size and shape rather than relying on fixed administrative boundaries or arbitrary hierarchies.
- A configurable, component‑based architecture that enables researchers to plug in alternative algorithms for each stage, facilitating comparative studies and future extensions.
- Empirical evidence that Voronoi‑based aggregation can achieve a better trade‑off between privacy (k‑anonymity) and data utility (geographic precision, low suppression) than existing methods.
Limitations noted by the authors include the current focus on two‑dimensional Euclidean space, which may not capture real‑world constraints such as road networks, natural barriers, or multi‑layer GIS data. Voronoi cells can also be irregular, potentially complicating downstream analyses that assume regular shapes. The system does not yet incorporate differential privacy guarantees or handle streaming data.
Future work suggested involves extending VBAS to three‑dimensional or network‑constrained spaces, integrating real‑time data streams, coupling the framework with differential privacy mechanisms, and adding shape‑regularization constraints to produce more analytically convenient regions.
Overall, the paper presents a compelling approach that leverages computational geometry to improve the balance between privacy protection and geographic utility in health‑care data releases, offering a flexible platform for both practitioners and researchers to tailor anonymization strategies to their specific data characteristics and analytical needs.
Comments & Academic Discussion
Loading comments...
Leave a Comment