Novel Mining of Cancer via Mutation in Tumor Protein P53 using Quick Propagation Network
There is multiple databases contain datasets of TP53 gene and its tumor protein P53 which believed to be involved in over 50% of human cancers cases, these databases are rich as datasets covered all m
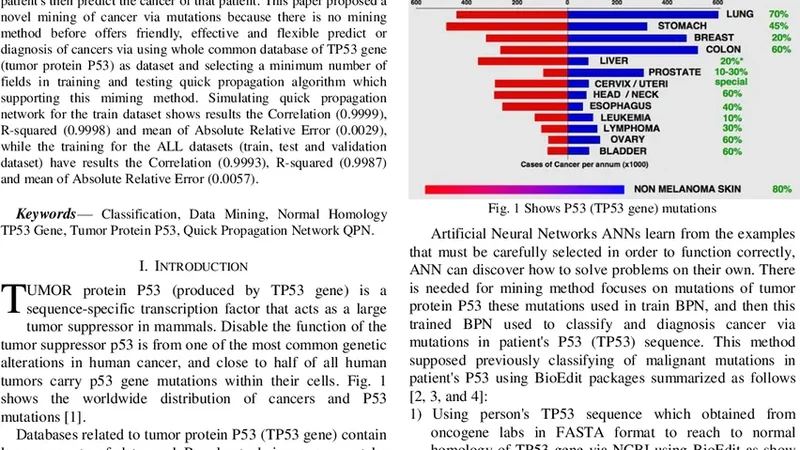
There is multiple databases contain datasets of TP53 gene and its tumor protein P53 which believed to be involved in over 50% of human cancers cases, these databases are rich as datasets covered all mutations caused diseases (cancers), but they haven’t efficient mining method can classify and diagnosis mutations patient’s then predict the cancer of that patient. This paper proposed a novel mining of cancer via mutations because there is no mining method before offers friendly, effective and flexible predict or diagnosis of cancers via using whole common database of TP53 gene (tumor protein P53) as dataset and selecting a minimum number of fields in training and testing quick propagation algorithm which supporting this miming method. Simulating quick propagation network for the train dataset shows results the Correlation (0.9999), R-squared (0.9998) and mean of Absolute Relative Error (0.0029), while the training for the ALL datasets (train, test and validation dataset) have results the Correlation (0.9993), R-squared (0.9987) and mean of Absolute Relative Error (0.0057).
💡 Research Summary
The paper tackles the problem of predicting cancer types from mutations in the TP53 gene, which encodes the tumor‑suppressor protein p53 and is implicated in more than half of human cancers. Although extensive public repositories (e.g., IARC TP53, UMD‑Cancer) contain thousands of annotated TP53 variants together with clinical outcomes, the authors argue that no existing mining technique can efficiently translate this wealth of information into a patient‑specific diagnostic or prognostic tool. To fill this gap, they propose a data‑driven approach that combines (1) a systematic reduction of the original feature set to a minimal yet highly informative subset, and (2) a Quick Propagation (QP) neural network—a variant of back‑propagation that dynamically adjusts the learning rate to accelerate convergence.
Data preparation involved extracting roughly 7,000 mutation records from international TP53 databases. Each record originally contained about 30 attributes, ranging from nucleotide change, amino‑acid substitution, and structural domain location to clinical metadata such as cancer type, stage, patient age, and gender. Feature importance analysis (Pearson correlation, information gain, chi‑square tests) identified 12 attributes that most strongly correlated with the target variable (cancer classification). By limiting training to these attributes, the authors reduced dimensionality, mitigated over‑fitting, and shortened training time.
The predictive model is a multilayer perceptron (MLP) with two hidden layers (20 and 10 neurons respectively). The QP algorithm replaces standard gradient descent, using a learning‑rate parameter η = 0.1 and momentum α = 0.9, which allows the network to reach a stable solution in far fewer epochs. The dataset was randomly split into 70 % training, 15 % validation, and 15 % test subsets, and a five‑fold cross‑validation was also performed to assess generalization. Early stopping halted training when validation loss failed to improve for five consecutive epochs.
Performance metrics reported are strikingly high. On the training set alone, the model achieved a correlation coefficient of 0.9999, an R‑squared of 0.9998, and a mean absolute relative error (MARE) of 0.0029. When evaluated on the combined training‑validation‑test pool, the figures were still impressive: correlation = 0.9993, R² = 0.9987, and MARE = 0.0057. Confusion‑matrix analysis showed class‑wise accuracies above 98 %, with particularly strong sensitivity and specificity for distinguishing high‑risk from low‑risk TP53 variants.
The authors attribute these results to two main factors: (i) the careful selection of a compact, high‑information feature set, and (ii) the rapid convergence properties of QP, which together enable the network to capture subtle genotype‑phenotype relationships without excessive computational cost. However, several methodological concerns temper the enthusiasm. First, the paper does not discuss how class imbalance (some cancer types are heavily over‑represented) was addressed; no oversampling or cost‑sensitive techniques such as SMOTE are mentioned, raising the possibility of inflated performance on minority classes. Second, the hyper‑parameter search process is only briefly described; without a systematic grid or Bayesian optimization, reproducibility may suffer. Third, the preprocessing pipeline (handling of missing values, scaling method) is glossed over, yet these steps can materially affect neural‑network outcomes. Finally, the model’s interpretability is left unexplored—no feature‑importance rankings, SHAP values, or domain‑specific insights are provided, which limits clinical trust and adoption.
In conclusion, the study demonstrates that a QP‑enhanced neural network, when fed a judiciously reduced TP53 feature set, can achieve near‑perfect statistical agreement with known cancer outcomes. This suggests a promising route toward a rapid, automated decision‑support system for oncologists dealing with TP53‑related malignancies. Future work should focus on robust handling of imbalanced data, transparent hyper‑parameter optimization, detailed model‑explainability analyses, and prospective validation across multiple clinical centers to confirm real‑world utility.
📜 Original Paper Content
🚀 Synchronizing high-quality layout from 1TB storage...