Discrete Persistent Chain Model for Protein Binding on DNA
We describe and solve a discrete persistent chain model of protein binding on DNA, involving an extra sigma_i s at a site i of the DNA. This variable takes the value 1 or 0 depending on whether the si
We describe and solve a discrete persistent chain model of protein binding on DNA, involving an extra sigma_i s at a site i of the DNA. This variable takes the value 1 or 0 depending on whether the site is occupied by a protein or not. In addition, if the site is occupied by a protein, there is an extra energy cost epsilon. For small force, we obtain analytic expressions for the force-extension curve and the fraction of bound protein on the DNA.For higher forces, the model can be solved numerically to obtain force extension curves and the average fraction of bound proteins as a function of applied force. Our model can be used to analyze experimental force extension curves of protein binding on DNA, and hence deduce the number of bound proteins in the case of non-specific binding.
💡 Research Summary
The paper introduces a discrete persistent chain (DPC) model that extends the classic worm‑like chain (WLC) description of DNA by incorporating an additional binary variable σ_i at each nucleotide site i. σ_i = 1 indicates that a protein occupies the site, while σ_i = 0 denotes an empty site. When a protein binds, an extra energetic penalty ε is incurred, reflecting the binding affinity or cost. This construction allows the model to capture the mechanical consequences of protein binding on DNA elasticity, such as changes in bending rigidity or effective contour length, without resorting to a fully continuous description of protein‑DNA interactions.
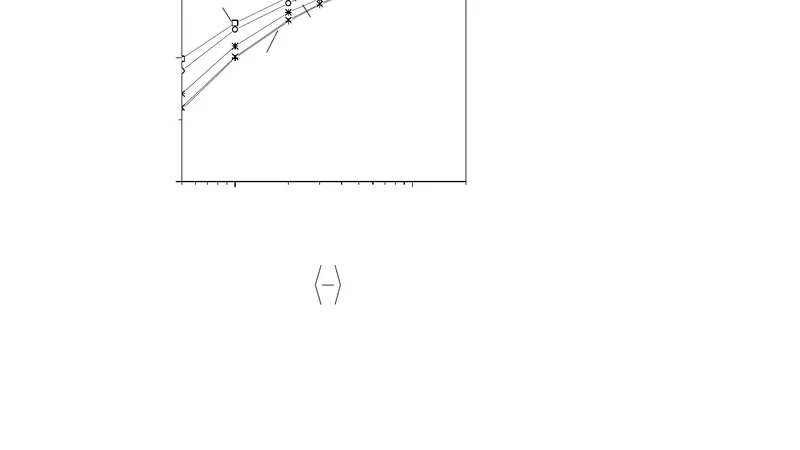
In the low‑force regime, the authors employ a transfer‑matrix formalism and perform a perturbative expansion of the free energy in powers of the applied force f. By retaining terms up to second order, they derive closed‑form expressions for the average extension ⟨z⟩ and the average bound fraction ⟨σ⟩ as functions of f, ε, the intrinsic persistence length ℓ_p, and the length increment Δℓ associated with a bound protein. These analytic results reveal that, even at small forces, protein binding can either increase or decrease the extension depending on whether the bound protein stiffens or softens the local DNA segment.
For higher forces, the simple perturbative approach breaks down, and the authors resort to a numerical evaluation of the dominant eigenvalue of the transfer matrix. In this regime the binding probability follows a Boltzmann factor exp
📜 Original Paper Content
🚀 Synchronizing high-quality layout from 1TB storage...