Affine and Regional Dynamic Time Warpng
Pointwise matches between two time series are of great importance in time series analysis, and dynamic time warping (DTW) is known to provide generally reasonable matches. There are situations where time series alignment should be invariant to scaling and offset in amplitude or where local regions of the considered time series should be strongly reflected in pointwise matches. Two different variants of DTW, affine DTW (ADTW) and regional DTW (RDTW), are proposed to handle scaling and offset in amplitude and provide regional emphasis respectively. Furthermore, ADTW and RDTW can be combined in two different ways to generate alignments that incorporate advantages from both methods, where the affine model can be applied either globally to the entire time series or locally to each region. The proposed alignment methods outperform DTW on specific simulated datasets, and one-nearest-neighbor classifiers using their associated difference measures are competitive with the difference measures associated with state-of-the-art alignment methods on real datasets.
💡 Research Summary
Dynamic Time Warping (DTW) is a widely used algorithm for aligning two time‑series that may be out of phase due to non‑linear temporal variations. Although DTW often yields reasonable alignments, it suffers from two important shortcomings: (1) it does not account for global amplitude scaling and offset between the series, and (2) it treats every point equally, ignoring the fact that certain local regions may be more informative for a particular application. To address these issues, the authors introduce two complementary extensions of DTW—Affine DTW (ADTW) and Regional DTW (RDTW)—and then combine them in two hybrid schemes, Global‑Affine RDTW (GARDTW) and Local‑Affine RDTW (LARDTW).
ADTW models one series as a scaled (c) and offset (e) version of the other while allowing arbitrary temporal warping. The optimal alignment path p, together with the global scaling and offset, is obtained by a hard Expectation‑Maximization (EM) loop. In the E‑step, DTW is applied to the transformed series (c·t+e) to update p; in the M‑step, closed‑form least‑squares formulas compute the new (c, e) given the current path. This alternating procedure guarantees a monotonic decrease of the objective and converges in a few iterations. The computational cost is O(n·nc·wb), where n is the series length, wb the Sakoe‑Chiba band width, and nc the number of EM iterations, while memory usage remains O(n·wb).
RDTW replaces the pointwise distance d(s_a, t_b) with a regional distance dr(s_a, t_b, wh) that averages the squared differences over a symmetric window of half‑width wh around each matched pair. By pre‑computing cumulative sums, most entries of the dynamic‑programming table can be updated in O(1) time, preserving the overall O(n·wb) complexity of DTW. The window width controls the emphasis on local structures; a small wh focuses on fine‑grained details, whereas a larger wh smooths over noise.
GARDTW integrates ADTW and RDTW at the global level: a single scaling and offset are estimated for the whole series, but the alignment cost uses the regional distance dr. The same hard EM scheme is employed, with the E‑step performing RDTW on the currently transformed series and the M‑step updating (c, e) via convex optimization. LARDTW goes further by allowing each matched region to have its own scaling and offset. For every pair (s_a, t_b) the optimal local parameters (c_{a,b}, e_{a,b}) are derived analytically from the data inside the window, and the overall alignment is found by standard DTW dynamic programming using these locally adapted distances.
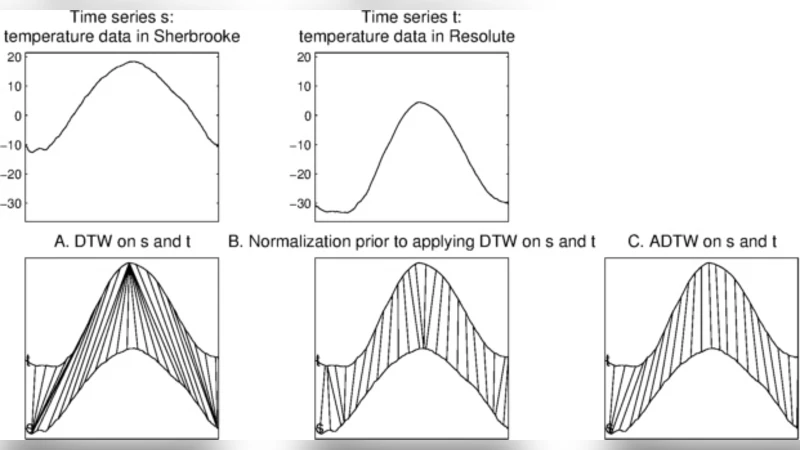
The authors evaluate the four methods on synthetic and real data. In a temperature‑series experiment, where two yearly temperature curves differ by a global scaling and offset, ADTW and GARDTW correctly align the peaks, whereas plain DTW produces pathological many‑to‑one matches. In a biomedical example with motor unit potentials composed of overlapping muscle‑fiber potentials, RDTW and LARDTW successfully separate the overlapping components, while DTW merges them incorrectly. Finally, on 85 datasets from the UCR Time Series Archive, 1‑Nearest‑Neighbour classification using the proposed distance measures achieves average accuracy improvements of 1–2 percentage points over standard DTW, Soft‑DTW, and Shape‑DTW. Parameter selection (scaling bounds, window width) is performed via cross‑validation, and the code is released publicly.
In summary, the paper presents a coherent framework that simultaneously handles amplitude scaling/offset and regional importance in time‑series alignment. The hard EM approach provides a practical trade‑off between computational efficiency and solution quality, and the experimental results demonstrate that the new distances are competitive with, and often superior to, state‑of‑the‑art alignment methods across diverse domains such as climate, biomedical signals, and general pattern recognition.
Comments & Academic Discussion
Loading comments...
Leave a Comment