The Time Machine: A Simulation Approach for Stochastic Trees
In the following paper we consider a simulation technique for stochastic trees. One of the most important areas in computational genetics is the calculation and subsequent maximization of the likeliho
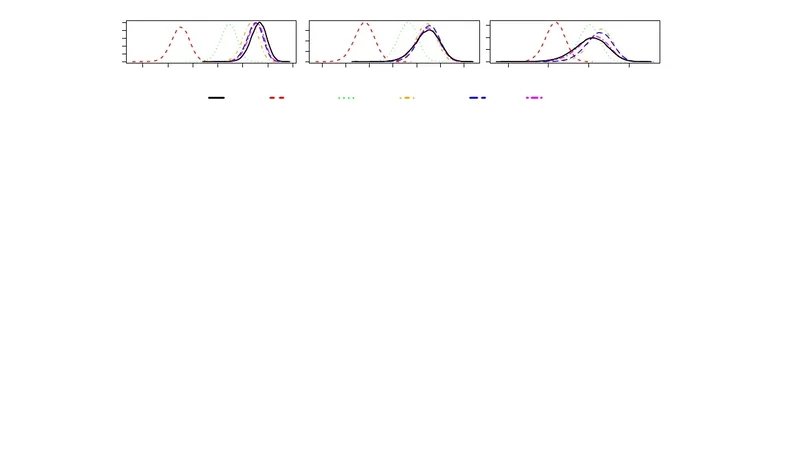
In the following paper we consider a simulation technique for stochastic trees. One of the most important areas in computational genetics is the calculation and subsequent maximization of the likelihood function associated to such models. This typically consists of using importance sampling (IS) and sequential Monte Carlo (SMC) techniques. The approach proceeds by simulating the tree, backward in time from observed data, to a most recent common ancestor (MRCA). However, in many cases, the computational time and variance of estimators are often too high to make standard approaches useful. In this paper we propose to stop the simulation, subsequently yielding biased estimates of the likelihood surface. The bias is investigated from a theoretical point of view. Results from simulation studies are also given to investigate the balance between loss of accuracy, saving in computing time and variance reduction.
💡 Research Summary
The paper addresses a fundamental computational bottleneck in the likelihood evaluation of stochastic tree models, which are central to many problems in computational genetics such as coalescent inference and phylogenetic reconstruction. Traditional importance‑sampling (IS) and sequential Monte‑Carlo (SMC) methods generate genealogical trees by simulating backwards from the observed genetic data to the most recent common ancestor (MRCA). While theoretically unbiased, these approaches become prohibitively expensive as the depth of the tree increases, and the variance of the resulting likelihood estimator often explodes because the importance weights can become extremely imbalanced.
To mitigate both the computational cost and the variance, the authors propose a deliberately biased technique: they stop the backward simulation before reaching the MRCA and replace the unfinished portion of the tree with a tractable approximation based on the prior distribution and a simple conditional likelihood model. This “simulation‑stop” strategy introduces a systematic bias, but the authors argue that the bias can be quantified, bounded, and, crucially, outweighed by the gains in speed and variance reduction.
The theoretical contribution begins with a decomposition of the full likelihood (L) into two components: the contribution from the simulated segment up to a stopping time (\tau) and the contribution from the un‑simulated remainder. The simulated segment is handled exactly as in standard IS/SMC, preserving the usual importance weights. For the remainder, the authors replace the exact transition kernel with a surrogate kernel (q) that is easy to sample from and analytically tractable. They derive an explicit expression for the bias (\Delta = \mathbb{E}
📜 Original Paper Content
🚀 Synchronizing high-quality layout from 1TB storage...