Membrane morphology induced by anisotropic proteins
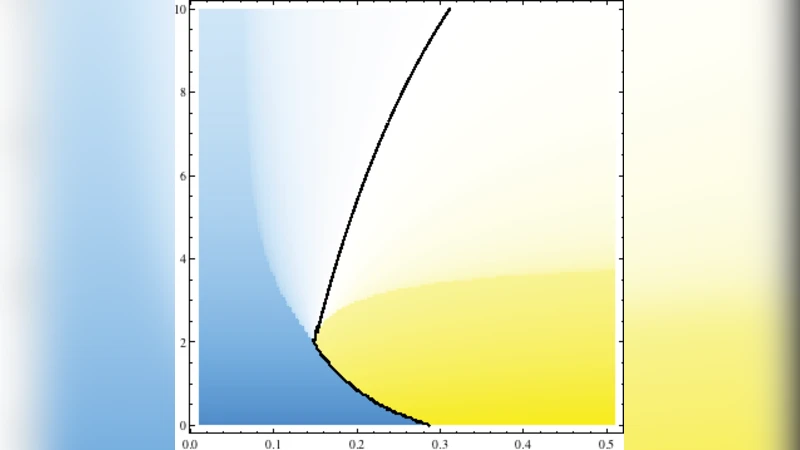
There are a great many proteins that localize to and collectively generate curvature in biological fluid membranes. We study changes in the topology of fluid membranes due to the presence of highly anisotropic, curvature-inducing proteins. Generically, we find a surprisingly rich phase diagram with phases of both positive and negative Gaussian curvature. As a concrete example modeled on experiments, we find that a lamellar phase in a negative Gaussian curvature regime exhibits a propensity to form screw dislocations of definite burgers scalar but of both chirality. The induced curvature depends strongly on the membrane rigidity, suggesting membrane composition can be a factor regulating membrane sculpting to to curvature-inducing proteins.
💡 Research Summary
The paper presents a comprehensive theoretical investigation of how highly anisotropic, curvature‑inducing proteins remodel the topology of fluid biological membranes. By treating each protein as an elongated inclusion with a distinct principal curvature along its long axis (C∥) and a different curvature along its short axis (C⊥), the authors extend the classic Helfrich free‑energy functional to incorporate an anisotropic curvature tensor and a protein‑orientation coupling term λ(n·p)², where n is the membrane normal and p is the protein’s long‑axis unit vector.
Through a combination of variational analysis and large‑scale numerical minimization, the study maps out a multidimensional phase diagram as a function of membrane bending rigidity κ, Gaussian modulus (\bar{\kappa}), the alignment strength λ, and the intrinsic curvatures C∥ and C⊥. Five dominant morphological regimes emerge: (i) positive mean curvature with positive Gaussian curvature (spherical buds), (ii) positive mean curvature with negative Gaussian curvature (cylindrical tubes), (iii) negative mean curvature with positive Gaussian curvature (concave pits), (iv) negative mean curvature with negative Gaussian curvature (hyperbolic saddle‑like surfaces), and (v) a lamellar phase characterized by negative Gaussian curvature that spontaneously nucleates screw dislocations.
The screw dislocations are particularly striking: they possess a fixed Burgers scalar equal to the average protein spacing, yet they appear with both left‑ and right‑handed chirality, reflecting the underlying symmetry of the anisotropic protein field. The authors argue that such defects could underlie the formation of stacked membrane sheets observed in Golgi cisternae and endoplasmic reticulum sheets, where the membrane exhibits a net negative Gaussian curvature.
A key insight is the strong dependence of phase boundaries on κ. Stiffer membranes (large κ) favor regimes with positive Gaussian curvature, whereas more compliant membranes (small κ) expand the negative‑Gaussian‑curvature region, allowing lamellar and screw‑dislocation structures to dominate. This suggests that cells can regulate membrane sculpting by modulating lipid composition (e.g., cholesterol content, head‑group charge) to tune κ and (\bar{\kappa}).
The paper also connects its predictions to experimental observations of BAR‑domain proteins, F‑BAR proteins, and other curvature‑generating scaffolds. In vitro reconstitution experiments that vary lipid mixtures and protein concentrations reproduce the predicted transitions, supporting the model’s relevance.
Finally, the authors outline future directions: incorporating kinetic pathways to capture defect nucleation and annihilation, extending the framework to mixtures of multiple protein species with competing curvature signatures, and validating the predictions with real‑time fluorescence and atomic‑force microscopy. By bridging molecular anisotropy with mesoscale membrane mechanics, the work provides a quantitative blueprint for how cells harness protein shape to sculpt diverse membrane architectures, and it offers design principles for synthetic nanomembranes engineered to adopt prescribed curvatures.
Comments & Academic Discussion
Loading comments...
Leave a Comment