Role of anisotropy for protein-protein encounter
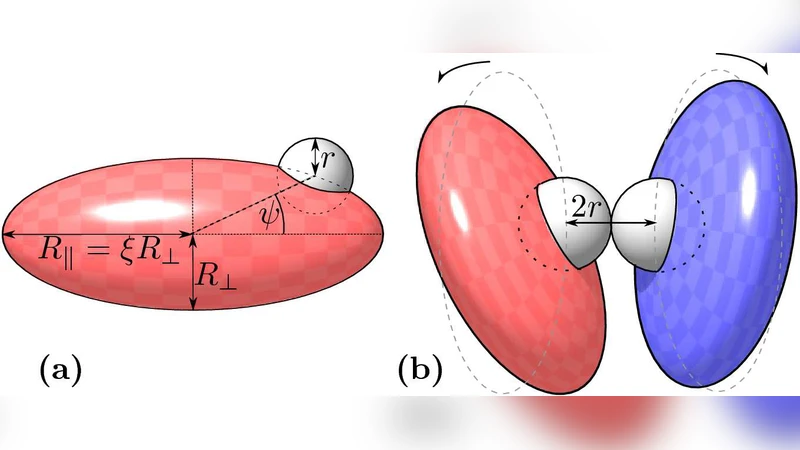
Protein-protein interactions comprise both transport and reaction steps. During the transport step, anisotropy of proteins and their complexes is important both for hydrodynamic diffusion and accessibility of the binding site. Using a Brownian dynamics approach and extensive computer simulations, we quantify the effect of anisotropy on the encounter rate of ellipsoidal particles covered with spherical encounter patches. We show that the encounter rate $k$ depends on the aspect ratios $\xi$ mainly through steric effects, while anisotropic diffusion has only a little effect. Calculating analytically the crossover times from anisotropic to isotropic diffusion in three dimensions, we find that they are much smaller than typical protein encounter times, in agreement with our numerical results.
💡 Research Summary
The paper investigates how the anisotropic shape of proteins influences the rate at which two proteins encounter each other during the diffusion‑controlled transport phase of a protein‑protein interaction. The authors model proteins as rigid ellipsoids decorated with spherical “encounter patches” that represent the localized binding sites. By varying the aspect ratio ξ (the ratio of the long to short axis) from unity (a sphere) up to five, they generate a systematic series of particle geometries that span a realistic range of protein anisotropy.
A Brownian dynamics framework is employed to simulate the stochastic motion of many independent particle pairs in three‑dimensional space. The simulations incorporate only hard‑core steric repulsion, deliberately excluding electrostatic or specific attractive forces, so that any observed changes in the encounter rate k can be attributed solely to geometric factors. For each ξ, thousands of trajectories are generated, and the mean first‑passage time to a successful encounter (i.e., simultaneous contact of the two patches) is recorded.
The results reveal two distinct contributions to the encounter kinetics. First, a strong steric effect emerges: as ξ increases, the ellipsoid becomes more elongated, reducing the angular window through which a patch can approach another particle without colliding with the body of the partner. Consequently, the effective “capture cross‑section” shrinks, and the measured encounter rate drops by a factor of roughly two to six across the explored aspect‑ratio range. This steric reduction dominates the ξ‑dependence of k.
Second, the authors examine the role of anisotropic diffusion. Because an ellipsoid has direction‑dependent translational and rotational diffusion coefficients, one might expect that the initial, orientation‑biased motion could significantly affect encounter probabilities. To quantify this, they analytically calculate the crossover time τ at which the anisotropic diffusion regime gives way to an effectively isotropic diffusion regime. The expression for τ depends on particle size, solvent viscosity, temperature, and the aspect ratio, and yields values on the order of 10⁻⁸ seconds for the simulated systems. This timescale is orders of magnitude shorter than typical protein‑protein encounter times (microseconds to milliseconds). Direct comparison with the Brownian dynamics data confirms that, after a few nanoseconds, the motion of the ellipsoids is already well described by isotropic diffusion, and the correction factor f_dif(ξ) associated with anisotropic diffusion remains essentially unity.
Putting the two effects together, the authors propose a compact phenomenological formula:
k(ξ) ≈ k₀ · f_steric(ξ) · f_dif(ξ),
where k₀ is the encounter rate for a spherical particle, f_steric(ξ) captures the reduction in capture cross‑section caused by elongation, and f_dif(ξ) accounts for any residual anisotropic diffusion influence. Their numerical data show that f_dif(ξ) ≈ 1 for all ξ, confirming that the steric term alone explains the observed variation in k.
The study has several important implications. It validates the widespread practice of using spherical approximations in coarse‑grained models of protein association when the primary interest lies in overall encounter rates, provided that the steric accessibility of binding sites is correctly represented. At the same time, it highlights that the geometric anisotropy of a protein can dramatically modulate its effective capture radius, a factor that should be taken into account in rational drug design, protein engineering, and kinetic modeling of cellular signaling pathways. Moreover, the analytical estimate of τ offers a useful benchmark for deciding when a full anisotropic diffusion treatment is necessary; for most biologically relevant systems, the rapid crossover to isotropic behavior justifies the use of simpler diffusion kernels.
Future extensions could incorporate electrostatic steering, flexible linkers, or multiple patches to explore how additional physical interactions interplay with shape anisotropy. Nevertheless, the present work provides a clear, quantitative foundation for understanding and modeling the impact of protein shape on the fundamental encounter step that precedes binding.
Comments & Academic Discussion
Loading comments...
Leave a Comment