Protein-mediated DNA Loop Formation and Breakdown in a Fluctuating Environment
Living cells provide a fluctuating, out-of-equilibrium environment in which genes must coordinate cellular function. DNA looping, which is a common means of regulating transcription, is very much a stochastic process; the loops arise from the thermal motion of the DNA and other fluctuations of the cellular environment. We present single-molecule measurements of DNA loop formation and breakdown when an artificial fluctuating force, applied to mimic a fluctuating cellular environment, is imposed on the DNA. We show that loop formation is greatly enhanced in the presence of noise of only a fraction of $k_B T$, yet find that hypothetical regulatory schemes that employ mechanical tension in the DNA–as a sensitive switch to control transcription–can be surprisingly robust due to a fortuitous cancellation of noise effects.
💡 Research Summary
The authors set out to quantify how the intrinsically fluctuating physical environment inside living cells influences the stochastic formation and breakdown of protein‑mediated DNA loops, a key mechanism for transcriptional regulation. Using a single‑molecule assay that combines optical tweezers with a magnetic‑bead torque system, they tethered a 1‑kilobase DNA fragment containing two LacI operator sites between two beads. By applying a static tensile force ranging from 0 to 2 pN and superimposing a high‑frequency (≈10 kHz) white‑noise component, they created an artificial “fluctuating force” whose root‑mean‑square amplitude corresponds to only 0.1–0.5 kBT—comparable to the magnitude of intracellular mechanical noise generated by motor proteins and cytoskeletal dynamics.
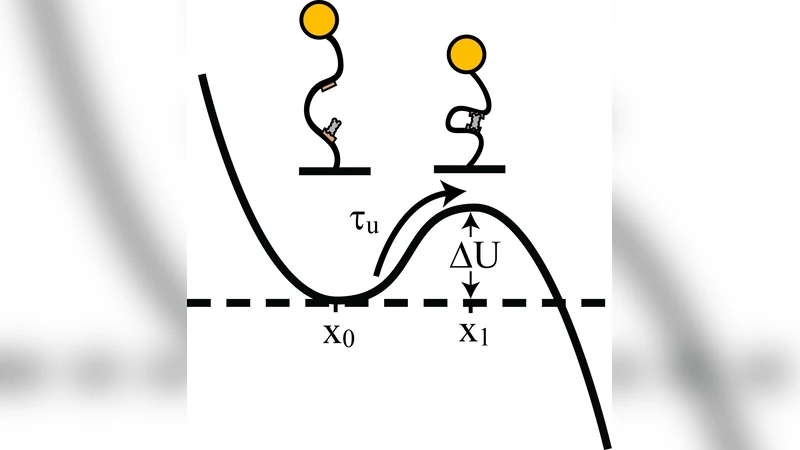
In the absence of noise, the expected tension‑dependent behavior was observed: increasing static force reduced the probability of loop formation and lengthened the average waiting time before a loop formed, reflecting the well‑known tension‑induced suppression of DNA looping. When the same static forces were combined with the small‑amplitude noise, however, loop formation rates (k_on) increased dramatically—up to fourfold at low tensions (<0.5 pN). Loop breakdown rates (k_off) were largely unchanged, showing a slight decrease at higher noise amplitudes. The authors interpret this asymmetry as a consequence of noise‑induced transient reductions in the bending free energy of the DNA, which raises the instantaneous probability that the two operator sites come into proximity, thereby accelerating loop capture without substantially affecting loop stability.
To assess the functional relevance of these findings for gene regulation, the team modeled a hypothetical transcriptional switch in which mechanical tension on the DNA acts as a binary control: below a critical force (~1 pN) loops form readily, turning transcription “on,” whereas above this threshold loops are disfavored, turning transcription “off.” Incorporating the experimentally measured noise‑enhanced k_on and modestly reduced k_off into the model revealed a fortuitous cancellation: the steepness of the force‑response curve and the location of the switching threshold remained essentially unchanged despite the presence of noise. Consequently, a regulatory scheme that relies on mechanical tension as a switch would be robust against the modest fluctuations typical of the intracellular milieu.
These results have two major implications. First, they provide direct experimental evidence that cellular mechanical noise can actively promote DNA looping, suggesting that transcriptional networks operate in a dynamic, non‑equilibrium regime where stochastic physical forces contribute to regulatory outcomes. Second, they demonstrate that tension‑based regulatory mechanisms can be intrinsically buffered against such noise, offering a plausible design principle by which cells achieve rapid, reversible control of gene expression without sacrificing reliability.
Methodologically, the study establishes a versatile platform for probing other DNA‑protein interactions under controlled noisy conditions. By varying the noise spectrum, DNA length, or bending rigidity, one could mimic diverse intracellular contexts, from chromatin remodeling to transcription factor binding under crowding. Future work could integrate real‑time force traces from motor proteins (e.g., myosin, kinesin) to generate biologically realistic “bio‑synthetic” noise profiles, thereby refining our understanding of how mechanical fluctuations shape the behavior of genetic circuits in living cells.
Comments & Academic Discussion
Loading comments...
Leave a Comment