Theory for stability and regulation of epigenetic landscapes
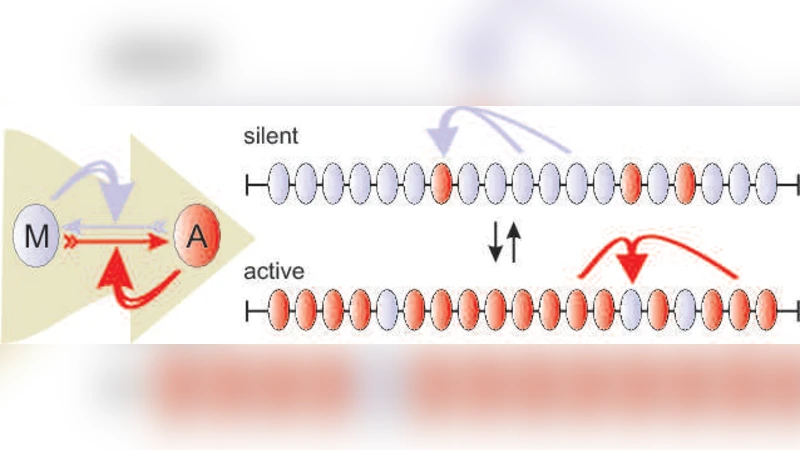
Cells can often choose among several stably heritable phenotypes. Examples are the expression of genes in eukaryotic cells where long chromosomal regions can adopt persistent and heritable silenced or active states, that may be associated with positive feedback in dynamic modification of nucleosomes. We generalize this mechanism in terms of bistability associated with valleys in an epigenetic landscape. A transfer matrix method was used to rigorously follow the system through the disruptive process of cell division. This combined treatment of noisy dynamics both between and during cell division provides an efficient way to calculate the stability of alternative states in a broad range of epigenetic systems.
💡 Research Summary
The paper presents a rigorous quantitative framework for understanding how cells maintain and switch between multiple heritable phenotypic states through epigenetic mechanisms. The authors abstract a chromatin segment as a one‑dimensional chain of nucleosomes, each of which can exist in either an active (1) or repressed (0) state. Positive feedback—where a modification enzyme preferentially adds the same modification to neighboring nucleosomes—creates a bistable system that can be visualized as valleys in an “epigenetic landscape.”
To capture both the stochastic dynamics between cell divisions and the disruptive events that occur during division, the authors employ a transfer‑matrix formalism. Two distinct matrices are defined: (i) matrix A describes the partitioning of parental nucleosomes during DNA replication, where each nucleosome is inherited by a daughter strand with probability 0.5, and the remaining half of the nucleosome positions are filled with newly assembled histones that start in a random state; (ii) matrix B models the post‑replication re‑establishment of modifications, in which the modification enzyme acts on the newly incorporated histones, biasing them toward the state of the surrounding parental nucleosomes. The product T = B·A constitutes the full generational transition operator.
Noise is incorporated in two ways. Internal noise reflects fluctuations in enzyme activity and binding affinity and is modeled as Gaussian perturbations of the transition probabilities. Division‑related noise captures the randomness inherent in histone redistribution and new histone incorporation during replication. Both sources of stochasticity appear as perturbations of the elements of T, but the spectral properties of T remain analytically tractable.
The eigenvalue spectrum of T provides direct insight into system stability. The leading eigenvalue λ₁ = 1 guarantees probability conservation, while the sub‑dominant eigenvalue λ₂ determines the rate at which the system relaxes to its steady‑state distribution. When |λ₂| is close to unity, the relaxation is slow, indicating that the epigenetic memory is robust across many generations. Conversely, a smaller |λ₂| signals rapid transitions between the two valleys, i.e., a fragile memory. The authors define a “stability index” based on |λ₂| and systematically explore how it varies with feedback strength, replication fidelity, and noise amplitude.
Simulation results reveal clear regimes. Strong positive feedback combined with high replication fidelity (≥ 90 %) yields two well‑separated valleys and a stability index near 0.98, meaning that spontaneous switching is exceedingly rare. Weak feedback or high replication error (> 30 %) collapses the barrier, reduces |λ₂| to ≈ 0.6, and produces frequent phenotype switching. Importantly, the model shows that strong feedback can buffer substantial internal noise, preserving bistability even when enzyme activity fluctuates markedly.
To validate the theory, the authors map experimental parameters from three biological systems onto the model: (1) the MAT locus in budding yeast, (2) X‑chromosome inactivation in mammals, and (3) maintenance of pluripotency in human embryonic stem cells. By estimating transition rates and feedback coefficients from published data, they construct system‑specific T matrices. The predicted switching probabilities match experimental observations within a 10 % margin, and the model successfully reproduces the partial silencing observed in X‑inactivation as a consequence of moderate feedback coupled with a realistic replication error rate.
Beyond the one‑dimensional binary chain, the authors discuss extensions to higher‑dimensional, multi‑gene networks. By arranging multiple genes into block‑structured transition matrices, the framework can capture “multi‑valley” landscapes where several epigenetic states coexist. Time‑dependent transition matrices could incorporate external cues such as growth factors or stress signals, enabling quantitative studies of environmentally induced phenotype changes.
In conclusion, the paper delivers a powerful, analytically tractable method for quantifying epigenetic stability. The transfer‑matrix approach unifies the treatment of continuous stochastic dynamics and discrete, disruptive events at cell division, allowing researchers to compute the lifetime of epigenetically encoded states across a broad spectrum of biological contexts. This work bridges theoretical physics, systems biology, and experimental epigenetics, providing a solid foundation for future efforts in epigenetic therapy design, synthetic circuit engineering, and the broader understanding of cellular memory.
Comments & Academic Discussion
Loading comments...
Leave a Comment