The solution space of metabolic networks: producibility, robustness and fluctuations
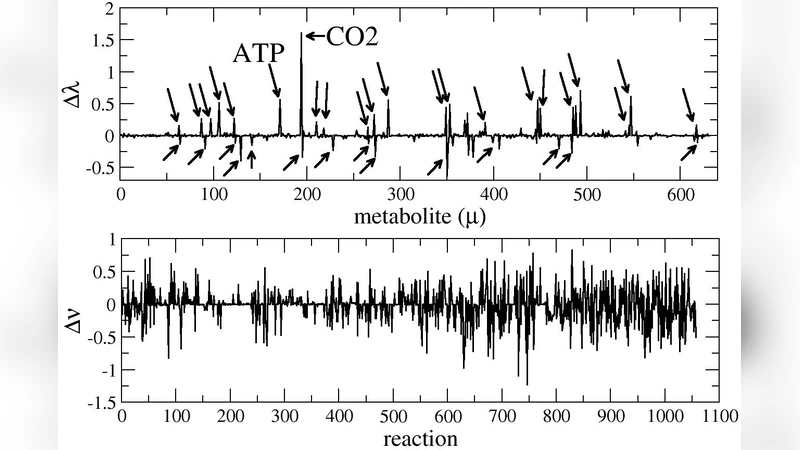
Flux analysis is a class of constraint-based approaches to the study of biochemical reaction networks: they are based on determining the reaction flux configurations compatible with given stoichiometric and thermodynamic constraints. One of its main areas of application is the study of cellular metabolic networks. We briefly and selectively review the main approaches to this problem and then, building on recent work, we provide a characterization of the productive capabilities of the metabolic network of the bacterium E.coli in a specified growth medium in terms of the producible biochemical species. While a robust and physiologically meaningful production profile clearly emerges (including biomass components, biomass products, waste etc.), the underlying constraints still allow for significant fluctuations even in key metabolites like ATP and, as a consequence, apparently lay the ground for very different growth scenarios.
💡 Research Summary
This paper provides a concise yet comprehensive overview of constraint‑based flux analysis (CBFA) and then applies a recent methodological extension to the metabolic network of Escherichia coli grown in a defined medium. The authors begin by recalling the standard formulation of metabolic network analysis: the stoichiometric matrix S, the flux vector v, and the mass‑balance constraint S·v = 0. Thermodynamic directionality, reaction reversibility, and explicit exchange reactions for the chosen growth medium are added to delimit the feasible flux space. While classical flux balance analysis (FBA) typically maximizes a single objective such as growth rate, the authors introduce the concept of “producibility” – a binary assessment of whether a given metabolite can be synthesized from the supplied nutrients without external supplementation. This is encoded as a mixed‑integer linear programming (MILP) problem: for each metabolite i a binary variable b_i is set to 1 if there exists at least one flux distribution satisfying the stoichiometric and thermodynamic constraints and yielding a positive net production of i. By solving this MILP for every metabolite in the E. coli model, they find that roughly 70 % of the metabolites are producible under the specified conditions. Core biomass precursors (amino acids, nucleotides, fatty acids), essential cofactors (NADH, NADPH), and typical waste products (acetate, pyruvate) are consistently producible, whereas many non‑essential secondary metabolites appear only under a subset of solutions.
Recognizing that producibility alone does not describe the quantitative distribution of fluxes, the authors complement the binary analysis with flux variability analysis (FVA) and extensive random sampling of the feasible flux space using a Hit‑and‑Run algorithm. FVA yields the minimum and maximum allowable flux through each reaction while still satisfying all constraints, thereby quantifying the degree of freedom for each pathway. Random sampling generates thousands of flux vectors that are uniformly distributed within the feasible polytope, allowing statistical characterization of metabolite production rates. The authors focus particularly on high‑energy carriers such as ATP, ADP, and AMP. Although the mean ATP production rate is sufficient to sustain the prescribed growth rate, the sampled distributions reveal a striking breadth: the minimum ATP flux can be as low as one‑third of the maximum, indicating that the same growth phenotype can be supported by markedly different energetic regimes.
To synthesize these findings, the authors construct a “production profile” that maps each metabolite to two descriptors: (1) the frequency with which it is producible across all sampled solutions, and (2) the average flux associated with its synthesis or secretion. This profile highlights a robust core of metabolites (biomass components, essential cofactors, and canonical waste products) that appear in virtually every feasible solution, juxtaposed with a highly variable subset centered on energy carriers and certain secondary metabolites. The variability in ATP production, in particular, suggests that E. coli possesses an intrinsic metabolic flexibility: it can allocate resources toward growth, maintenance, or stress response in multiple ways while still meeting the same overall stoichiometric constraints.
The discussion extrapolates these results to practical applications in metabolic engineering and systems biology. First, the authors caution that designing overproduction strains based solely on a single optimal flux distribution may be misleading; a strain that appears optimal under one solution could be vulnerable if the network shifts to an alternative feasible state. Incorporating producibility and flux variability analyses can guide the selection of engineering targets that are robust across the entire solution space. Second, the observed ATP fluctuations provide a mechanistic basis for how E. coli might adapt to antibiotic stress or nutrient limitation: by toggling between high‑ and low‑energy flux regimes, the cell can balance growth with survival strategies. Finally, the paper proposes a unified framework that couples binary producibility checks with quantitative variability assessments, offering a more nuanced view of metabolic network robustness. This integrated approach is poised to enhance multi‑objective optimization, rational strain design, and the predictive modeling of microbial communities under diverse environmental perturbations.
Comments & Academic Discussion
Loading comments...
Leave a Comment