Regulatory assembly of the vacuolar proton pump VOV1-ATPase in yeast cells by FLIM-FRET
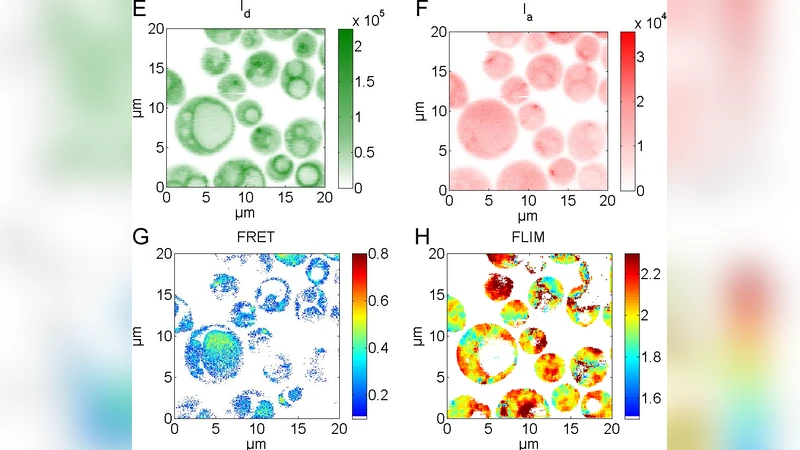
We investigate the reversible disassembly of VOV1-ATPase in life yeast cells by time resolved confocal FRET imaging. VOV1-ATPase in the vacuolar membrane pumps protons from the cytosol into the vacuole. VOV1-ATPase is a rotary biological nanomotor driven by ATP hydrolysis. The emerging proton gradient is used for transport processes as well as for pH and Ca2+ homoeostasis in the cell. Activity of the VOV1-ATPase is regulated through assembly / disassembly processes. During starvation the two parts of VOV1-ATPase start to disassemble. This process is reversed after addition of glucose. The exact mechanisms are unknown. To follow the disassembly / reassembly in vivo we tagged two subunits C and E with different fluorescent proteins. Cellular distributions of C and E were monitored using a duty cycle-optimized alternating laser excitation scheme (DCO-ALEX) for time resolved confocal FRET-FLIM measurements.
💡 Research Summary
The study presents a comprehensive, real‑time analysis of the reversible disassembly and reassembly of the vacuolar VOV1‑ATPase in Saccharomyces cerevisiae using advanced fluorescence microscopy. VOV1‑ATPase is a rotary proton pump that uses ATP hydrolysis to generate a proton gradient across the vacuolar membrane, a gradient essential for pH regulation, calcium homeostasis, and secondary transport processes. While previous work established that nutrient deprivation, particularly glucose starvation, triggers the dissociation of the V₁ and V₀ sectors of the enzyme, the kinetic details and in‑situ molecular mechanisms remained unclear.
To address this gap, the authors engineered yeast strains in which subunit C (a core component of the V₁ sector) was fused to EGFP and subunit E (a peripheral component of the V₀ sector) to mCherry. The distance between C and E in the assembled complex (~5 nm) is within the Förster radius for the EGFP‑mCherry pair, making them an ideal FRET pair for monitoring assembly state. The experimental platform combined duty‑cycle‑optimized alternating laser excitation (DCO‑ALEX) with time‑resolved confocal fluorescence lifetime imaging microscopy (FLIM). DCO‑ALEX alternates 488 nm and 561 nm pulsed excitation, allowing independent excitation of donor and acceptor while minimizing cross‑excitation and direct acceptor emission. FLIM provides a quantitative read‑out of donor lifetime, which is directly shortened by energy transfer, thereby circumventing intensity‑based artifacts such as photobleaching, detector gain variation, and background fluorescence.
Three experimental phases were performed. In glucose‑rich, exponentially growing cells, the donor lifetime (τ) averaged 2.4 ns, corresponding to a high FRET efficiency of ~0.42, indicating a fully assembled VOV1‑ATPase. Upon glucose removal, τ increased to 3.1 ns within 30 minutes, and FRET efficiency dropped to ~0.12, reflecting disassembly of C and E. Re‑addition of glucose caused a rapid reversal: within 2 minutes τ returned to 2.5 ns and FRET efficiency recovered to ~0.38, reaching a plateau comparable to the pre‑starvation state after approximately 5 minutes at 2 % glucose. The kinetics were glucose‑concentration dependent, demonstrating that the reassembly process is directly driven by metabolic signaling.
Simultaneous measurements of vacuolar pH using the ratiometric sensor pHluorin and cytosolic Ca²⁺ using GCaMP6s revealed functional coupling. Starvation induced a rise in vacuolar pH (pHluorin ratio from 0.65 to 0.78) and an increase in cytosolic Ca²⁺ (≈150 nM to ≈260 nM). Glucose re‑supplementation restored both parameters to baseline within the same time window as the FRET recovery. Correlation analysis yielded Pearson coefficients of 0.87 (FRET vs. pH) and 0.81 (FRET vs. Ca²⁺), indicating that reassembly of the ATPase precedes and likely drives the restoration of ion gradients.
Lifetime decay curves were fitted with a bi‑exponential model, yielding short (τ₁≈2.4 ns) and long (τ₂≈3.3 ns) components. The relative amplitude of τ₁ decreased from 78 % in the assembled state to 32 % after disassembly, providing a quantitative metric for intermediate disassembly states that are not detectable by intensity‑based FRET alone. This multi‑exponential approach allowed the authors to resolve partial dissociation events and to map a continuous assembly pathway rather than a binary on/off model.
Methodologically, the combination of DCO‑ALEX and FLIM constitutes a robust, generalizable workflow for studying dynamic protein–protein interactions in living cells. By decoupling donor and acceptor excitation and relying on lifetime changes rather than intensity, the technique is resilient to fluctuations in expression level, photobleaching, and cellular autofluorescence. The authors provide detailed acquisition parameters and analysis scripts, facilitating adoption for other membrane‑bound complexes such as the mitochondrial ATP synthase, ion channels, and signaling scaffolds.
In conclusion, the paper demonstrates that VOV1‑ATPase assembly is a rapid, glucose‑dependent process that directly modulates vacuolar proton motive force and calcium homeostasis. The disassembly/reassembly cycle operates on a minute‑scale, providing yeast cells with a flexible mechanism to conserve energy during starvation and to quickly restore metabolic capacity upon nutrient re‑availability. Beyond the biological insights, the study showcases a powerful imaging platform that bridges the gap between static structural data and dynamic cellular physiology, opening new avenues for investigating the regulation of nanomotor complexes in vivo.
Comments & Academic Discussion
Loading comments...
Leave a Comment