A systematically coarse-grained model for DNA, and its predictions for persistence length, stacking, twist, and chirality
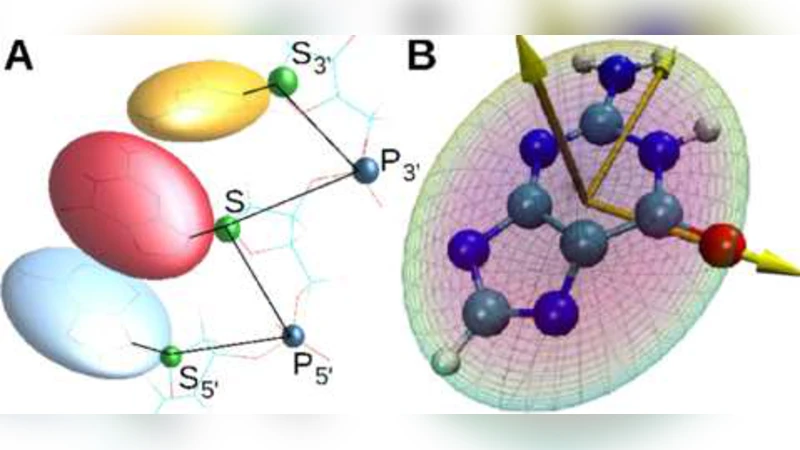
We introduce a coarse-grained model of DNA with bases modeled as rigid-body ellipsoids to capture their anisotropic stereochemistry. Interaction potentials are all physicochemical and generated from all-atom simulation/parameterization with minimal phenomenology. Persistence length, degree of stacking, and twist are studied by molecular dynamics simulation as functions of temperature, salt concentration, sequence, interaction potential strength, and local position along the chain, for both single- and double-stranded DNA where appropriate. The model of DNA shows several phase transitions and crossover regimes in addition to dehybridization, including unstacking, untwisting, and collapse which affect mechanical properties such as rigidity and persistence length. The model also exhibits chirality with a stable right-handed and metastable left-handed helix.
💡 Research Summary
The authors present a physically grounded coarse‑grained model of DNA in which each nucleotide base is represented as a rigid‑body ellipsoid, thereby preserving the anisotropic stereochemistry that governs stacking and hydrogen‑bonding interactions. All interaction potentials are derived from all‑atom molecular dynamics simulations and parameterized with minimal phenomenological input, incorporating electrostatic screening, solvent‑mediated forces, and ion effects directly from the underlying atomistic data. This approach enables the model to respond naturally to changes in temperature, salt concentration, sequence composition, and interaction strength without ad‑hoc adjustments.
Molecular dynamics simulations were performed on both single‑stranded and double‑stranded DNA to evaluate three key mechanical observables: persistence length, degree of base stacking, and helical twist angle. The results reveal a rich landscape of phase‑like transitions. As temperature rises, stacking interactions weaken, leading to an “unstacking” transition that sharply reduces persistence length and concomitantly diminishes the twist angle—a process the authors term “untwisting.” Increasing ionic strength initially stabilizes the double helix by screening repulsive phosphate charges, thereby extending the persistence length. However, beyond a critical salt concentration the chain collapses into a compact state, indicating a separate “collapse” regime. Sequence dependence is evident: GC‑rich segments retain higher stacking energies and longer persistence lengths across a broad temperature range, whereas AT‑rich regions are more susceptible to thermal destabilization.
A particularly notable outcome is the emergence of chirality. The model spontaneously adopts a right‑handed helix as the global energy minimum, consistent with native B‑DNA. Under specific initial conditions, a left‑handed helix can persist as a metastable state, suggesting that DNA’s handedness can be toggled by external perturbations such as torque or ionic conditions. This dual‑handedness behavior provides a mechanistic framework for understanding rare left‑handed DNA forms observed experimentally.
Overall, the study demonstrates that a rigorously parameterized coarse‑grained representation can capture the interplay between stacking, electrostatics, and mechanical rigidity, reproducing known DNA behaviors while predicting new transition regimes. The model’s predictive power makes it a valuable tool for nanobiotechnology applications, DNA nanostructure design, and fundamental biophysical investigations where atomistic simulations are computationally prohibitive. Future work will focus on extending the model to longer genomic fragments, incorporating explicit protein‑DNA interactions, and validating the predicted transitions against single‑molecule experiments.
Comments & Academic Discussion
Loading comments...
Leave a Comment