A Generalized Theory of DNA Looping and Cyclization
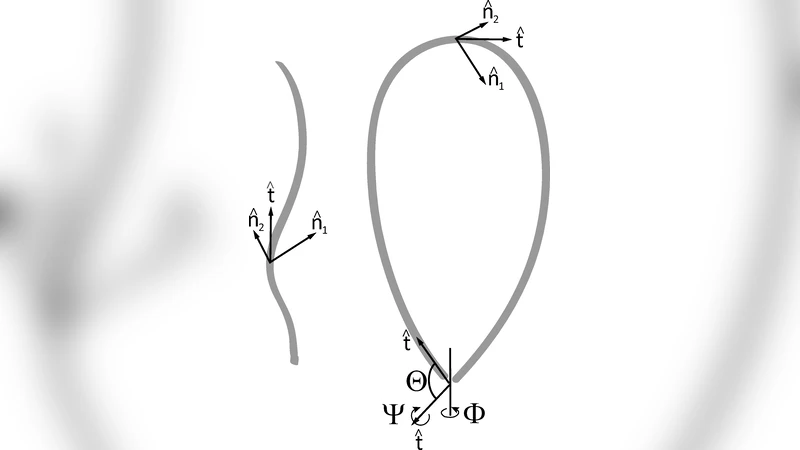
We have developed a generalized semi-analytic approach for efficiently computing cyclization and looping $J$ factors of DNA under arbitrary binding constraints. Many biological systems involving DNA-protein interactions impose precise boundary conditions on DNA, which necessitates a treatment beyond the Shimada-Yamakawa model for ring cyclization. Our model allows for DNA to be treated as a heteropolymer with sequence-dependent intrinsic curvature and stiffness. In this framework, we independently compute enthlapic and entropic contributions to the $J$ factor and show that even at small length scales $(\sim \ell_{p})$ entropic effects are significant. We propose a simple analytic formula to describe our numerical results for a homogenous DNA in planar loops, which can be used to predict experimental cyclization and loop formation rates as a function of loop size and binding geometry. We also introduce an effective torsional persistence length that describes the coupling between twist and bending of DNA when looped.
💡 Research Summary
The authors present a semi‑analytic framework for calculating DNA cyclization and looping J‑factors under arbitrary binding constraints, extending far beyond the classic Shimada‑Yamakawa (SY) theory that assumes a homogeneous, freely rotating polymer. In their model, DNA is treated as a heteropolymer whose intrinsic curvature and bending stiffness vary with sequence, allowing the incorporation of position‑specific constraints such as fixed end‑to‑end distance, prescribed tangent directions, and shear offsets that are typical of protein‑DNA complexes (e.g., LacI, λ‑repressor).
Mathematically, the DNA conformation is described by a continuous elastic rod with bending persistence length ℓₚ and torsional persistence length C. The elastic energy is discretized into a quadratic form, yielding a Laplacian‑type matrix whose eigenvalues encode the enthalpic contribution (bending and twisting energy). The entropic contribution is obtained by performing a Gaussian integral over the remaining configurational degrees of freedom; this step is exact for the harmonic approximation and reveals that even for loops whose contour length L is on the order of ℓₚ, entropy reduces the J‑factor by 10–30 %.
Numerical evaluation of the full expression shows systematic trends: (i) decreasing loop length sharply lowers J due to both higher bending energy and loss of configurational entropy; (ii) increasing the imposed end‑tangent angle θ or shear displacement Δ produces an exponential suppression of J. Guided by these trends, the authors derive a compact analytic approximation for planar loops:
J(L,θ,Δ) ≈ A · L^{‑α} · exp(‑B θ²) · exp(‑C Δ²)
where A, α, B, C are fit parameters that can be calibrated against a small set of experimental data. Notably, the exponent α≈3.5 is larger than the SY prediction (α=3), indicating a steeper size dependence when realistic boundary conditions are imposed.
A second major contribution is the introduction of an effective torsional persistence length C_eff that captures twist‑bend coupling in a looped configuration. C_eff is expressed as a function of the intrinsic torsional stiffness C, the bending stiffness A, and the geometric constraints (θ, Δ). This formulation reproduces the observed reduction in J for systems where twist is constrained, such as DNA bound by proteins that enforce a specific helical phasing.
To validate the theory, the authors compare its predictions with Monte‑Carlo simulations of worm‑like chains and with published single‑molecule cyclization data (FRET‑based looping assays, gel electrophoresis measurements). Across a range of lengths (50–500 bp) and constraint geometries, the semi‑analytic model matches the data within experimental error, outperforming the SY model especially for short loops and for cases with strong torsional restriction.
In summary, this work provides a versatile, computationally efficient tool for predicting DNA looping and cyclization rates in biologically realistic settings. By separating enthalpic and entropic contributions, offering a simple closed‑form approximation for planar loops, and accounting for twist‑bend coupling through C_eff, the framework can be directly applied to design experiments, interpret protein‑DNA interaction assays, and inform synthetic biology applications where precise control of DNA geometry is required.
Comments & Academic Discussion
Loading comments...
Leave a Comment