Maximally informative pairwise interactions in networks
Several types of biological networks have recently been shown to be accurately described by a maximum entropy model with pairwise interactions, also known as the Ising model. Here we present an approach for finding the optimal mappings between input signals and network states that allow the network to convey the maximal information about input signals drawn from a given distribution. This mapping also produces a set of linear equations for calculating the optimal Ising model coupling constants, as well as geometric properties that indicate the applicability of the pairwise Ising model. We show that the optimal pairwise interactions are on average zero for Gaussian and uniformly distributed inputs, whereas they are non-zero for inputs approximating those in natural environments. These non-zero network interactions are predicted to increase in strength as the noise in the response functions of each network node increases. This approach also suggests ways for how interactions with unmeasured parts of the network can be inferred from the parameters of response functions for the measured network nodes.
💡 Research Summary
This paper tackles a fundamental question in quantitative biology: how should a network of binary units be wired so that it transmits the maximal amount of information about external stimuli drawn from a known probability distribution? Building on the observation that many biological systems—neural populations, gene‑regulatory circuits, and immune cell ensembles—are well described by a maximum‑entropy (Ising) model with only pairwise couplings, the authors develop a principled framework for deriving the optimal mapping from inputs to network states and for computing the corresponding optimal Ising parameters.
Model formulation
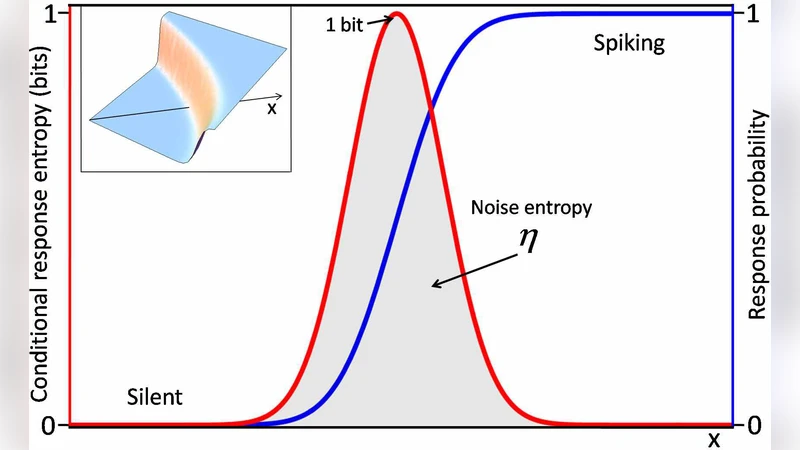
Each node i receives a continuous input vector x and produces a binary output σ_i∈{0,1}. The stochastic response is modeled by a sigmoidal transfer function
P(σ_i=1|x)=½
Comments & Academic Discussion
Loading comments...
Leave a Comment