Long delay times in reaction rates increase intrinsic fluctuations
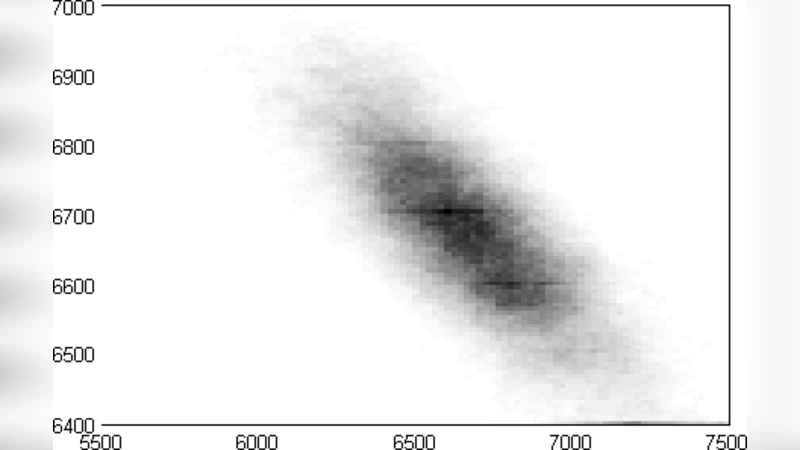
In spatially distributed cellular systems, it is often convenient to represent complicated auxiliary pathways and spatial transport by time-delayed reaction rates. Furthermore, many of the reactants appear in low numbers necessitating a probabilistic description. The coupling of delayed rates with stochastic dynamics leads to a probability conservation equation characterizing a non-Markovian process. A systematic approximation is derived that incorporates the effect of delayed rates on the characterization of molecular noise, valid in the limit of long delay time. By way of a simple example, we show that delayed reaction dynamics can only increase intrinsic fluctuations about the steady-state. The method is general enough to accommodate nonlinear transition rates, allowing characterization of fluctuations around a delay-induced limit cycle.
💡 Research Summary
The paper addresses a fundamental problem in stochastic modeling of spatially distributed cellular systems: how to incorporate auxiliary pathways and transport processes that are naturally delayed in time. Traditional stochastic chemical kinetics assumes Markovian dynamics, where reaction propensities depend only on the current state. When a reaction rate is delayed, the system becomes non‑Markovian because the propensity at time t depends on the state at an earlier time t − τ. This introduces a probability‑conserving master equation with a memory kernel, for which exact analytical solutions are rarely available, especially when molecule numbers are low and intrinsic noise dominates.
The authors first formulate the exact non‑Markovian master equation for a generic set of delayed reactions. They then focus on the limit of long delay times (τ ≫ characteristic relaxation time of the system). In this regime they develop a systematic approximation based on a Kramers‑Moyal expansion combined with a system‑size (Ω) expansion. The key result is that the deterministic mean‑field equations are unchanged by the delay – the average concentrations obey the same ordinary differential equations as in the delay‑free case – but the stochastic fluctuations acquire an additional positive contribution proportional to τ. In other words, the diffusion (variance) term in the corresponding Fokker‑Planck equation is enhanced, leading to larger intrinsic noise.
To illustrate the theory, the authors analyze two models. The first is a simple linear conversion reaction A → B with a delayed rate. By solving the approximate Fokker‑Planck equation they obtain an explicit expression for the steady‑state variance: σ² = σ₀² + k τ, where σ₀² is the variance without delay and k is the reaction rate constant. This demonstrates that any finite delay inevitably inflates the variance, regardless of the mean concentration.
The second example is a nonlinear system capable of generating a delay‑induced limit cycle via a Hopf bifurcation. Here the deterministic dynamics already exhibit oscillations, and the authors apply the same approximation to compute the covariance matrix around the periodic orbit. Their analysis shows that the amplitude of stochastic fluctuations around the limit cycle grows with τ, effectively broadening the distribution of phase and amplitude. This result is particularly relevant for biological oscillators such as circadian clocks or synthetic gene circuits, where delayed feedback is common and noise can have functional consequences.
The paper also discusses the validity range of the approximation. When τ is comparable to or shorter than the intrinsic timescales, higher‑order correction terms become important, and the first‑order long‑delay expansion loses accuracy. Nevertheless, the authors validate their theory with stochastic simulations using a delayed version of the Gillespie algorithm (Delayed Stochastic Simulation Algorithm, DSA). The simulations confirm that for τ an order of magnitude larger than the system’s relaxation time, the predicted variance increase matches the numerical results with high precision.
In summary, the study provides a clear, analytically tractable framework for quantifying how delayed reaction rates amplify intrinsic fluctuations in stochastic biochemical networks. By separating the deterministic mean‑field behavior from the stochastic correction, it offers a practical tool for researchers modeling gene regulation, signaling pathways, or any cellular process where transport or intermediate steps introduce significant time lags. The findings suggest that neglecting delays can lead to systematic underestimation of noise, which may affect the design and interpretation of experiments in synthetic biology and systems biology.
Comments & Academic Discussion
Loading comments...
Leave a Comment