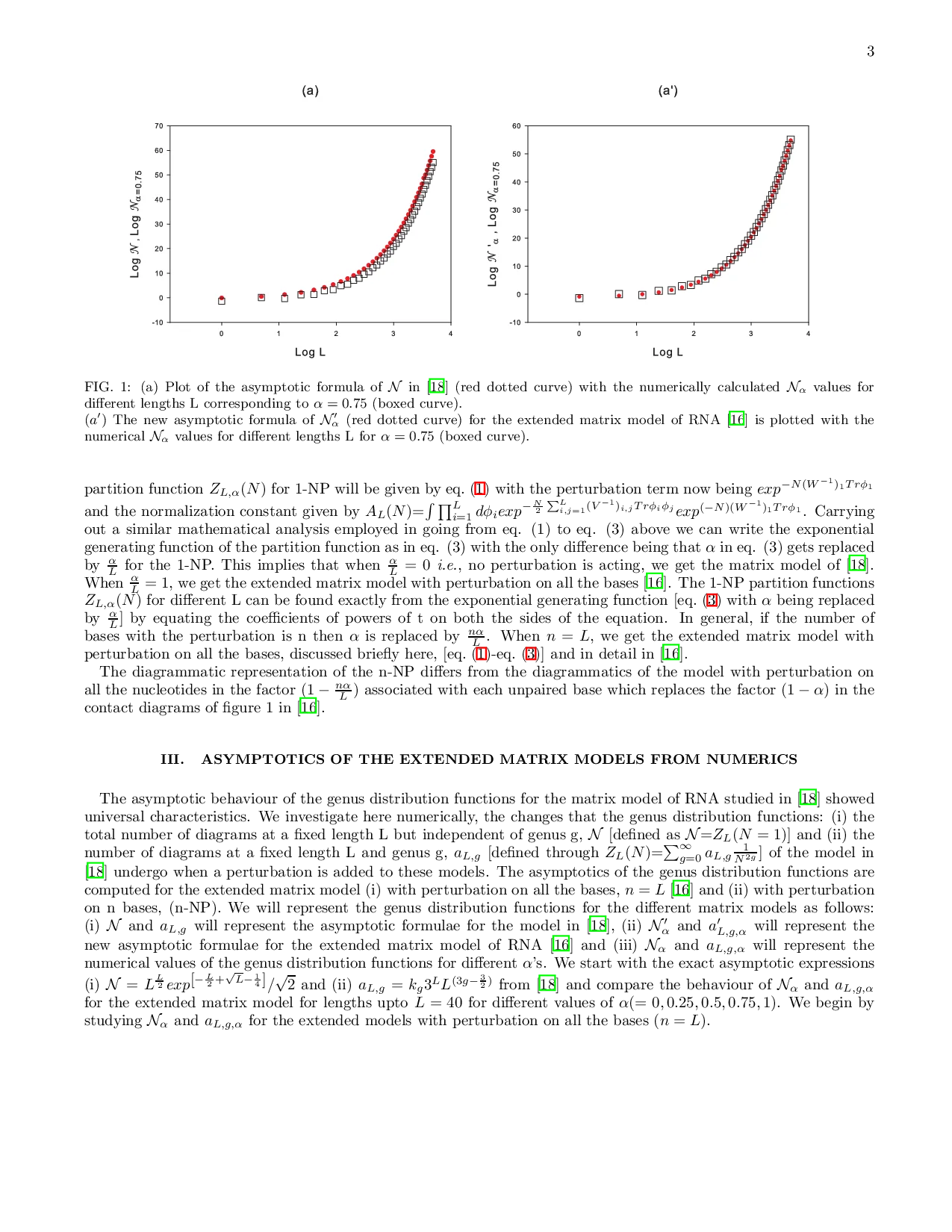
We study a matrix model of RNA in which an external perturbation acts on n nucleotides of the polymer chain. The effect of the perturbation appears in the exponential generating function of the partition function as a factor $(1-\frac{n\alpha}{L})$ [where $\alpha$ is the ratio of strengths of the original to the perturbed term and L is length of the chain]. The asymptotic behaviour of the genus distribution functions for the extended matrix model are analyzed numerically when (i) $n=L$ and (ii) $n=1$. In these matrix models of RNA, as $n\alpha/L$ is increased from 0 to 1, it is found that the universality of the number of diagrams $a_{L, g}$ at a fixed length L and genus g changes from $3^{L}$ to $(3-\frac{n\alpha}{L})^{L}$ ($2^{L}$ when $n\alpha/L=1$) and the asymptotic expression of the total number of diagrams $\cal N$ at a fixed length L but independent of genus g, changes in the factor $\exp^{\sqrt{L}}$ to $\exp^{(1-\frac{n\alpha}{L})\sqrt{L}}$ ($exp^{0}=1$ when $n\alpha/L=1$)
Deep Dive into RNA matrix models with external interactions and their asymptotic behaviour.
We study a matrix model of RNA in which an external perturbation acts on n nucleotides of the polymer chain. The effect of the perturbation appears in the exponential generating function of the partition function as a factor $(1-\frac{n\alpha}{L})$ [where $\alpha$ is the ratio of strengths of the original to the perturbed term and L is length of the chain]. The asymptotic behaviour of the genus distribution functions for the extended matrix model are analyzed numerically when (i) $n=L$ and (ii) $n=1$. In these matrix models of RNA, as $n\alpha/L$ is increased from 0 to 1, it is found that the universality of the number of diagrams $a_{L, g}$ at a fixed length L and genus g changes from $3^{L}$ to $(3-\frac{n\alpha}{L})^{L}$ ($2^{L}$ when $n\alpha/L=1$) and the asymptotic expression of the total number of diagrams $\cal N$ at a fixed length L but independent of genus g, changes in the factor $\exp^{\sqrt{L}}$ to $\exp^{(1-\frac{n\alpha}{L})\sqrt{L}}$ ($exp^{0}=1$ when $n\alpha/L=1$)
Improved understanding of the process of folding of RNA finds its ultimate use in the prediction of the fully folded, partially folded and completely unfolded structures under physiological conditions [1]. Under these conditions, unfolding is a very slow process as compared to folding in the presence of a force. Application of a force increases the unfolding rate and we can therefore get the unfolded structures from the folded ones ( [1] and references therein). Experimental techniques of force induced measurements have proved successful in probing properties related to different aspects of RNA folding and unfolding, domain unfolding in proteins, in polysachharides and nucleic acids ( [2] and references therein). Experiments have been performed on the double helixed DNAs to study their elastic and structural properties using electric field, hydrodynamic flow among other methods of force application ( [3] and references therein). The advent of AFM technique served as an important tool in the study of the basic underlying framework of molecular structural biology. Over the years, optical tweezers and AFM (atomic force microscopy) techniques have been employed to study the physical, elastic and structural properties of the biomolecules by recording their force extension curves (FECs) and studying the force dependent dynamics and folding landscapes of the molecules ( [4,5,6,7,8,9] and references therein). The conformations of biopolymers (DNA, RNA and proteins) which are otherwise not accesible from the conventional methods of measurements: NMR spectroscopy and X-ray crystallography, are possible with the use of AFMs. These conformations help in revealing the underlying mechanical framework of the biological systems ( [2] and references therein). Mechanical unfolding and refolding of single RNA has been studied using force-ramp, hopping and force-jump methods ( [10] and references therein). In mechanical unfolding experiments, it has been observed that at a critical value of the applied force, the hairpin structure toggles between the folded and the unfolded states [11,12,13]. In these experiments, ionic concentrations play an important role. Experiments of Bustamante et al [12,13] have shown that the denaturation of RNA by a constant force involves multiple trajectories (for RNA hairpins and Tetrahymena thermophila ribozyme) while undergoing a transition from the folded structure state to the unfolded state. These trajectories depend on the point at which the force is applied [1,14]. This diverseness in the folding-unfolding pathways is due to the rugged energy landscape of RNA (consisting of many minima). Controlled/monitored force loading and unloading rates can be used to manipulate the single molecules of RNA into either their native or misfolded pathways. Different force unloading rates in experiments on TAR RNA molecules showed different types of trajectories associated with particular refolding characteristics ( [15] and references therein).
We discuss here very briefly, a generalization of the extended random matrix model of RNA folding proposed in [16] where the external perturbation acts on a single nucleotide (n = 1) and on n nucleotides (n ≤ L) in the polymer chain (we will refer to the two RNA models as 1-NP RNA model, with NP being Nucleotide Perturbation and n-NP RNA model respectively). In [16], the external perturbation acted on all the nucleotides in the polymer chain (i.e., n = L, n is the number of bases on which the force is acting). We briskly outline the extended matrix model of [16] for completeness and understanding and follow it up with results and comparative discussion for the 1-NP and n-NP models. Further, we present a detailed numerical analysis for the asymptotics of the extended matrix model of RNA with perturbation on all the nucleotides in the polymer chain. The genus distribution functions: the total number of diagrams at a fixed length L but independent of genus g, N and the number of diagrams at a fixed length L and genus g, a L,g of the matrix model of RNA in [17,18] are found to change in the presence of the external perturbation. We extend our numerical asymptotic analysis to the n-NP RNA model as well.
We review here, the effect when a perturbation acts on all the nucleotides in the polymer chain (n = L) studied in [16]. The nucleotide-nucleotide interaction partition function of the polymer chain with a perturbation on all the bases is
where
)iT rφi is the perturbation term, V i,j is an (L×L) symmetric matrix containing information on the interactions between the L nucleotides at positions i and j in the polymer chain, φ i are L independent (N×N) hermitian matrices and the observable i (1 + φ i ) is an ordered product over φ i ’s. We consider V i,j = v and W i = w where v gives the strength of interaction between the nucleotides at positions i and j (in these models, interaction between any two nucleotides of the chain is considered the same and equal to v) and w g
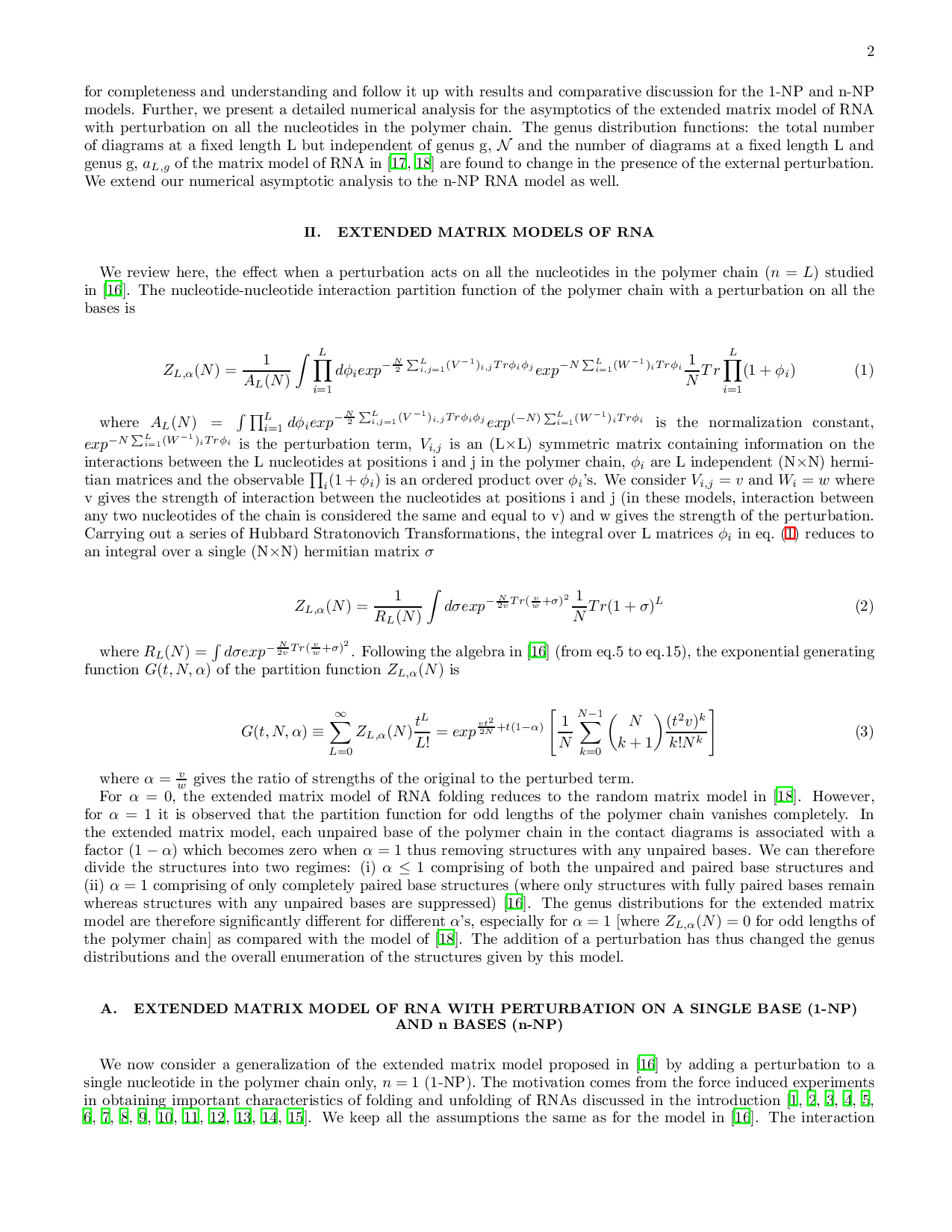
…(Full text truncated)…
This content is AI-processed based on ArXiv data.