On the accessibility of adaptive phenotypes of a bacterial metabolic network
The mechanisms by which adaptive phenotypes spread within an evolving population after their emergence are understood fairly well. Much less is known about the factors that influence the evolutionary accessibility of such phenotypes, a pre-requisite for their emergence in a population. Here, we investigate the influence of environmental quality on the accessibility of adaptive phenotypes of Escherichia coli’s central metabolic network. We used an established flux-balance model of metabolism as the basis for a genotype-phenotype map (GPM). We quantified the effects of seven qualitatively different environments (corresponding to both carbohydrate and gluconeogenic metabolic substrates) on the structure of this GPM. We found that the GPM has a more rugged structure in qualitatively poorer environments, suggesting that adaptive phenotypes could be intrinsically less accessible in such environments. Nevertheless, on average ~74% of the genotype can be altered by neutral drift, in the environment where the GPM is most rugged; this could allow evolving populations to circumvent such ruggedness. Furthermore, we found that the normalized mutual information (NMI) of genotype differences relative to phenotype differences, which measures the GPM’s capacity to transmit information about phenotype differences, is positively correlated with (simulation-based) estimates of the accessibility of adaptive phenotypes in different environments. These results are consistent with the predictions of a simple analytic theory and they suggest an intuitive information-theoretic principle for evolutionary adaptation; adaptation could be faster in environments where the GPM has a greater capacity to transmit information about phenotype differences.
💡 Research Summary
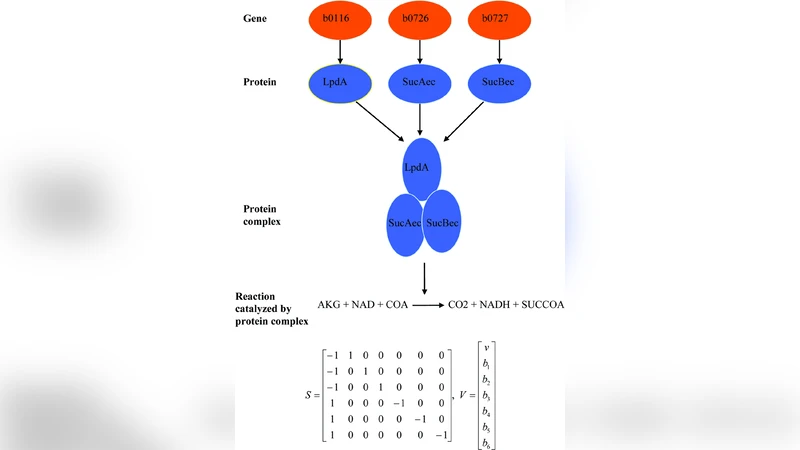
The paper addresses a gap in evolutionary biology: while the spread of adaptive phenotypes after they arise is relatively well understood, the factors that determine whether such phenotypes are evolutionarily accessible in the first place have received far less attention. To explore this, the authors focus on the central metabolic network of Escherichia coli and construct a genotype‑phenotype map (GPM) using a well‑established flux‑balance analysis (FBA) model. Each genotype is represented as a binary vector indicating the presence or absence of metabolic reactions, and the phenotype is defined as the maximal growth rate predicted by FBA under a given environmental condition.
Seven qualitatively distinct environments are examined, encompassing both carbohydrate substrates (glucose, lactose, maltose, etc.) and gluconeogenic precursors (pyruvate, malate, etc.). These environments span a gradient of “environmental quality,” from nutrient‑rich to nutrient‑poor. For each environment the authors sample a large number of genotypes, compute their growth rates, and thereby obtain a high‑dimensional mapping from genotype space to phenotype space.
The first major analysis quantifies the ruggedness of the GPM. Ruggedness is measured as the variance of phenotype differences among genotype pairs that are a fixed Hamming distance apart. The results show that poorer environments produce markedly more rugged GPMs: small genetic changes tend to cause large, unpredictable shifts in growth rate, implying that adaptive peaks are more isolated and that the landscape contains many “fitness traps.”
Despite this, the authors demonstrate that neutral drift remains a potent mechanism for traversing the landscape. By estimating the proportion of genotypes that can be altered by a single neutral mutation (i.e., a mutation that does not appreciably change growth rate), they find that even in the most rugged environment roughly 74 % of genotypes have at least one neutral neighbor. This high degree of neutrality suggests that evolving populations can sidestep rugged regions and continue to explore genotype space, eventually reaching adaptive phenotypes that would otherwise be inaccessible.
The second key finding links an information‑theoretic metric—normalized mutual information (NMI) between genotype differences and phenotype differences—to the empirical accessibility of adaptive phenotypes. NMI quantifies how much information about phenotypic change is conveyed by a given genetic change; higher NMI means that genotype‑phenotype mapping is more predictable. Simulations of adaptive walks in each environment reveal a positive correlation between NMI and the success rate of reaching adaptive phenotypes. In other words, environments that allow the GPM to transmit more information about phenotype differences also enable faster, more reliable adaptation.
To rationalize this correlation, the authors present a simple analytic model treating the GPM as a noisy communication channel. The model predicts that the channel capacity (captured by NMI) sets an upper bound on the speed of evolutionary search. Empirical data from the FBA‑based GPM conform to this prediction, supporting the authors’ proposal of an “information‑theoretic principle of adaptation”: evolutionary adaptation proceeds more quickly in environments where the genotype‑phenotype map has a greater capacity to convey information about phenotypic effects of mutations.
In summary, the study shows that environmental quality shapes the structural ruggedness of metabolic genotype‑phenotype maps, influencing the intrinsic accessibility of adaptive phenotypes. While poor environments generate more rugged landscapes, a high prevalence of neutral mutations and a strong genotype‑phenotype information flow can mitigate these constraints. By integrating flux‑balance modeling, landscape analysis, and information theory, the paper provides a comprehensive framework for understanding how environmental context governs both the possibility and the speed of evolutionary adaptation.