Precision analysis for standard deviation measurements of single fluorescent molecule images
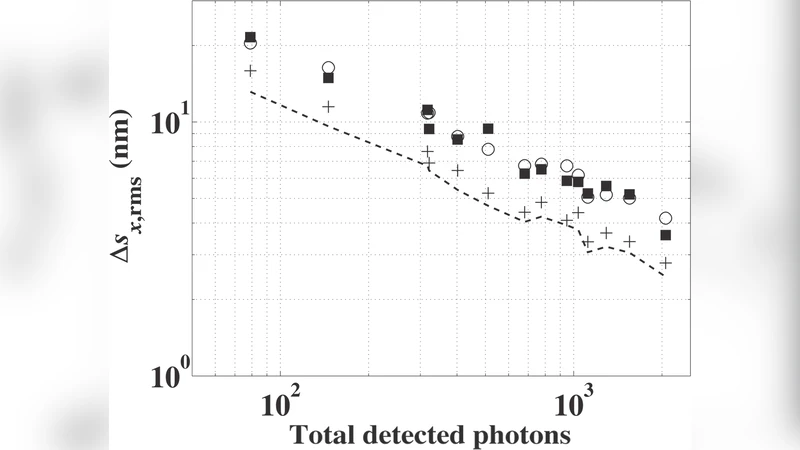
Standard deviation measurements of intensity profiles of stationary single fluorescent molecules are useful for studying axial localization, molecular orientation, and a fluorescence imaging system’s spatial resolution. Here we report on the analysis of the precision of standard deviation measurements of intensity profiles of single fluorescent molecules imaged using an EMCCD camera. We have developed an analytical expression for the standard deviation measurement error of a single image which is a function of the total number of detected photons, the background photon noise, and the camera pixel size. The theoretical results agree well with the experimental, simulation, and numerical integration results. Using this expression, we show that single-molecule standard deviation measurements offer nanometer precision for a large range of experimental parameters.
💡 Research Summary
The paper addresses the quantitative precision of measuring the standard deviation (σ) of intensity profiles from stationary single fluorescent molecules imaged with an EMCCD camera. The authors begin by modeling the image of a single molecule as a two‑dimensional Gaussian point spread function (PSF). Photon detection in each pixel is treated as a Poisson process, while the electron‑multiplying gain of the EMCCD introduces additional Gaussian read‑out noise with zero mean and variance σ_EM². The image intensity I(x, y) is parameterized by five variables: amplitude A, centroid coordinates (x₀, y₀), the Gaussian width σ, and a uniform background B. Using maximum‑likelihood estimation (MLE) for these parameters, the Fisher information matrix (FIM) is derived, leading to a Cramér‑Rao lower bound (CRLB) for the σ estimator. The resulting analytical expression for the standard‑deviation measurement error is
Δσ ≈ √
Comments & Academic Discussion
Loading comments...
Leave a Comment