Accuracy of direct gradient sensing by single cells
Many types of cells are able to accurately sense shallow gradients of chemicals across their diameters, allowing the cells to move towards or away from chemical sources. This chemotactic ability relies on the remarkable capacity of cells to infer gradients from particles randomly arriving at cell-surface receptors by diffusion. Whereas the physical limits of concentration sensing by cells have been explored, there is no theory for the physical limits of gradient sensing. Here, we derive such a theory, using as models a perfectly absorbing sphere and a perfectly monitoring sphere, which, respectively, infer gradients from the absorbed surface particle density or the positions of freely diffusing particles inside a spherical volume. We find that the perfectly absorbing sphere is superior to the perfectly monitoring sphere, both for concentration and gradient sensing, since previously observed particles are never remeasured. The superiority of the absorbing sphere helps explain the presence at the surfaces of cells of signal degrading enzymes, such as PDE for cAMP in Dictyostelium discoideum (Dicty) and BAR1 for mating factor alpha in Saccharomyces cerevisiae (budding yeast). Quantitatively, our theory compares favorably to recent measurements of Dicty moving up a cAMP gradient, suggesting these cells operate near the physical limits of gradient detection.
💡 Research Summary
The paper addresses a fundamental question in cellular chemotaxis: how accurately a single cell can infer a shallow chemical gradient from the stochastic arrival of ligand molecules by diffusion. While the physical limits of concentration sensing have been explored since Berg and Purcell’s classic work, a quantitative theory for gradient sensing was lacking. The authors fill this gap by constructing two idealized spherical models that capture the essential physics of ligand detection.
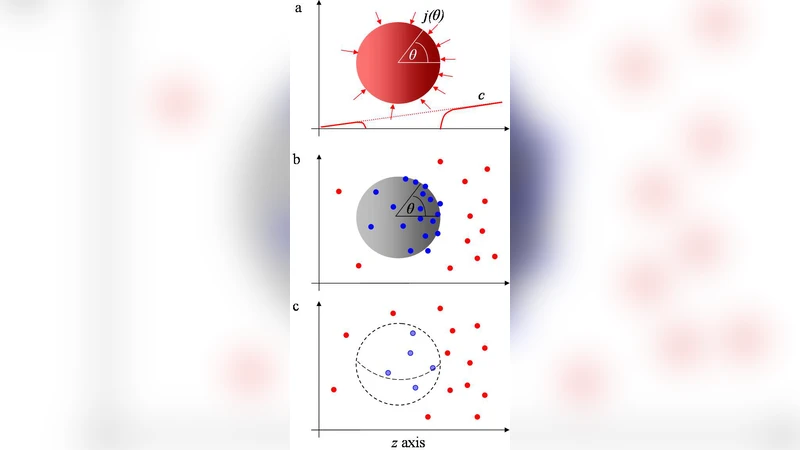
The first model, the “perfectly absorbing sphere,” assumes that any ligand that reaches the cell surface is instantly captured and removed, preventing any subsequent re‑measurement of the same molecule. The second model, the “perfectly monitoring sphere,” represents a cell that can monitor the positions of all ligands inside a spherical volume but does not remove them; consequently, a single ligand can be counted multiple times as it diffuses. Both models are treated analytically using diffusion equations and Poisson statistics to calculate the mean number of detected ligands, the variance of that number, and the variance of the difference between opposite sides of the sphere (the quantity that encodes the gradient).
For concentration sensing, the absorbing sphere yields a variance
\
Comments & Academic Discussion
Loading comments...
Leave a Comment