Polar Chemoreceptor Clustering by Coupled Trimers of Dimers
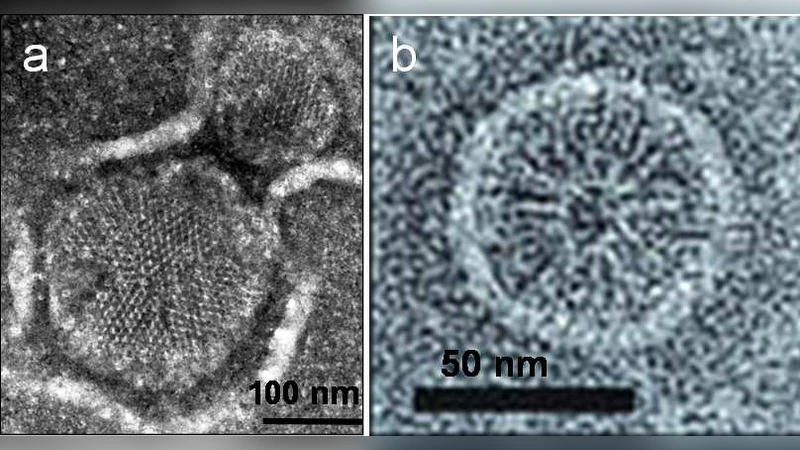
Receptors of bacterial chemotaxis form clusters at the cell poles, where clusters act as “antennas” to amplify small changes in ligand concentration. Interestingly, chemoreceptors cluster at multiple length scales. At the smallest scale, receptors form dimers, which assemble into stable timers of dimers. At a large scale, trimers form large polar clusters composed of thousands of receptors. Although much is known about the signaling properties emerging from receptor clusters, it is unknown how receptors localize at the cell poles and what the cluster-size determining factors are. Here, we present a model of polar receptor clustering based on coupled trimers of dimers, where cluster size is determined as a minimum of the cluster-membrane free energy. This energy has contributions from the cluster-membrane elastic energy, penalizing large clusters due to their high intrinsic curvature, and receptor-receptor coupling favoring large clusters. We find that the reduced cluster-membrane curvature mismatch at the curved cell poles leads to large and robust polar clusters in line with experimental observation, while lateral clusters are efficiently suppressed.
💡 Research Summary
The paper addresses a long‑standing question in bacterial chemotaxis: how chemoreceptors, which are known to form dimers and higher‑order trimers of dimers (TODs), become concentrated into massive polar clusters that function as highly sensitive “antennas.” While the structural organization of receptors into dimers and TODs is well documented, the physical principles that dictate the size, location, and stability of the resulting clusters have remained unclear. To fill this gap, the authors develop a quantitative model based on coupled trimers of dimers and determine the optimal cluster size by minimizing the total cluster‑membrane free energy.
The free‑energy functional comprises two antagonistic contributions. The first is an elastic penalty that arises because a receptor cluster has an intrinsic curvature (C₀) that may not match the curvature of the surrounding lipid bilayer (1/R). The mismatch Δκ = C₀ – 1/R generates an energy term proportional to N·(Δκ)², where N is the number of TODs in the cluster and α is an elastic constant reflecting membrane bending rigidity and receptor‑membrane coupling. The second contribution is a favorable receptor‑receptor coupling energy (γ) that scales with the number of pairwise contacts, i.e., –γ·N·(N‑1)/2. Adding a baseline insertion cost ε₀ per TOD yields the total free energy:
F(N) = N·ε₀ + α·N·(Δκ)² – γ·N·(N‑1)/2.
Minimizing F(N) with respect to N gives the equilibrium cluster size N*. Crucially, Δκ depends on the local membrane curvature. At the cell poles, the membrane is highly curved (R ≈ 0.5 µm for many rod‑shaped bacteria), which brings the membrane curvature close to the intrinsic curvature of the receptor complex. Consequently, the elastic penalty becomes small, allowing the attractive coupling term to dominate and drive the formation of very large clusters (N* in the thousands). In contrast, on the relatively flat lateral surface (R → ∞), Δκ is large, the elastic term overwhelms the coupling, and the free‑energy minimum occurs at very small N* (often 1–10), effectively suppressing lateral clusters.
Parameter values (ε₀ ≈ 2 k_BT, α ≈ 0.1 k_BT·nm², γ ≈ 0.05 k_BT) were estimated from structural data (TOD diameter ≈ 5 nm) and membrane mechanical measurements. Using these numbers, the model reproduces experimentally observed polar cluster sizes of 2,000–5,000 receptors and the near‑absence of sizable lateral aggregates. Monte‑Carlo simulations of the stochastic assembly process show a sharp, nucleation‑like transition at the poles: once a critical nucleus forms, rapid growth ensues because each additional TOD reduces the per‑unit free energy. On flat surfaces, the same nucleus is unstable and dissolves.
The authors further explore how ligand binding, which shifts receptors between active and inactive conformations, modulates γ. An increase in the active fraction raises the effective coupling, slightly enlarging polar clusters and enhancing signal amplification. However, the curvature‑dependent elastic term remains the dominant size‑determining factor, explaining why polar clustering is robust across a wide range of external stimuli.
In the discussion, the model is placed in the context of existing chemotaxis theories that focus on signaling dynamics (CheA activation, CheY phosphorylation) and on static structural arrangements (hexagonal lattices of TODs). The present work adds a missing physical layer: the interplay between membrane geometry and receptor‑receptor attraction. This insight predicts that any cellular process that alters pole curvature—such as cell division, shape mutants, or artificial curvature‑inducing scaffolds—should directly affect cluster size and, consequently, chemotactic sensitivity. The authors propose experimental tests using patterned substrates that impose defined curvatures or engineered receptor variants with altered intrinsic curvature.
In summary, the coupled‑trimers model provides a coherent, physics‑based explanation for three key observations: (1) the preferential polar localization of chemoreceptor clusters, (2) the emergence of very large, stable polar aggregates, and (3) the efficient suppression of lateral clusters. By showing that curvature mismatch and attractive coupling together set the free‑energy landscape, the study bridges membrane biophysics and signal transduction, offering a framework that can be extended to other membrane‑bound signaling complexes and to synthetic biology applications where spatial organization of receptors is desired.
Comments & Academic Discussion
Loading comments...
Leave a Comment