Non-specific DNA-protein interaction: Why proteins can diffuse along DNA
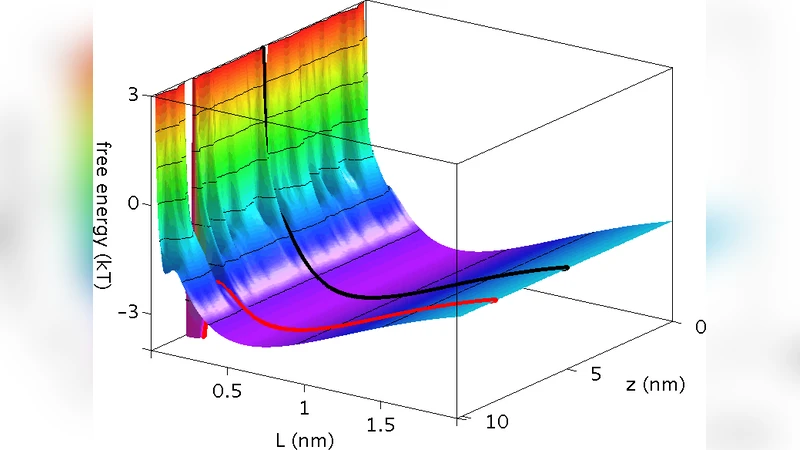
The structure of DNA Binding Proteins enables a strong interaction with their specific target site on DNA. However, recent single molecule experiment reported that proteins can diffuse on DNA. This suggests that the interactions between proteins and DNA play a role during the target search even far from the specific site. It is unclear how these non-specific interactions optimize the search process, and how the protein structure comes into play. Each nucleotide being negatively charged, one may think that the positive surface of DNA-BPs should electrostatically collapse onto DNA. Here we show by means of Monte Carlo simulations and analytical calculations that a counter-intuitive repulsion between the two oppositely charged macromolecules exists at a nanometer range. We also show that this repulsion is due to a local increase of the osmotic pressure exerted by the ions which are trapped at the interface. For the concave shape of DNA-BPs, and for realistic protein charge densities, we find that the repulsion pushes the protein in a free energy minimum at a distance from DNA. As a consequence, a favourable path exists along which proteins can slide without interacting with the DNA bases. When a protein encounters its target, the osmotic barrier is completely counter-balanced by the H-bond interaction, thus enabling the sequence recognition.
💡 Research Summary
The paper tackles a long‑standing paradox in the field of DNA‑binding proteins (DBPs): while the electrostatic picture predicts a strong attraction between the negatively charged DNA backbone and the positively charged surface of DBPs, single‑molecule experiments have repeatedly shown that many DBPs can slide along DNA without becoming tightly bound. To resolve this, the authors combine coarse‑grained Monte Carlo simulations with an analytical theory based on ion‑distribution and osmotic pressure. Their central finding is that, at nanometer separations (≈0.8–1.2 nm), a counter‑intuitive repulsive force emerges between the two oppositely charged macromolecules. This repulsion does not stem from direct electrostatic repulsion; rather, it originates from a local increase in osmotic pressure caused by ions that become trapped in the narrow gap between the DNA helix and the concave surface of the protein. The trapped ions raise the local ion concentration above that of the bulk solution, generating an osmotic pressure that pushes the protein away from the DNA surface.
When realistic protein charge densities (≈+5 e nm⁻²) and physiological salt concentrations (∼100 mM NaCl) are used, the free‑energy landscape exhibits a shallow minimum at a finite distance from the DNA. In this configuration the protein experiences a balance between the attractive electrostatic pull and the osmotic repulsion, allowing it to remain close enough to sense the DNA contour but far enough to avoid direct contact with the bases. Consequently, the protein can diffuse laterally along the DNA in a “sliding” mode that is essentially one‑dimensional and does not involve strong base‑specific interactions. This sliding pathway is energetically favorable because it avoids the high‑energy penalty associated with forcing the trapped ions out of the gap.
The model also explains how target recognition occurs. When the protein encounters its cognate sequence, specific hydrogen‑bonding interactions between amino‑acid side chains and DNA bases provide a large negative contribution to the free energy. This specific binding energy fully compensates the osmotic barrier, allowing the protein to collapse onto the DNA and form a stable, sequence‑specific complex. Thus, the non‑specific, osmotic‑driven repulsion governs the search phase, while the specific hydrogen‑bond network dominates the recognition phase.
By systematically varying protein curvature, charge density, and ionic strength, the authors demonstrate that the magnitude of the osmotic repulsion is highly sensitive to these parameters. Proteins with a more pronounced concave surface experience a stronger trapping of ions and therefore a deeper free‑energy well at a finite distance, whereas flatter proteins tend to be pulled directly onto the DNA. The work reconciles the “facilitated diffusion” concept (combining 1‑D sliding, hopping, and 3‑D diffusion) with a physically grounded mechanism that does not rely on ad‑hoc assumptions about weak binding.
In the discussion, the authors highlight several implications. First, the osmotic barrier provides a natural explanation for the observed diffusion coefficients of DBPs on DNA, which are often an order of magnitude slower than in free solution but still much faster than would be expected for a tightly bound complex. Second, the sensitivity of the barrier to ionic strength suggests that cells could regulate search efficiency by modulating local salt concentrations. Third, the insights could guide the engineering of synthetic DNA‑binding domains or CRISPR‑Cas variants with tunable search dynamics, simply by adjusting surface charge patterns or curvature.
Overall, the study presents a coherent, quantitatively validated picture: non‑specific DNA‑protein interactions are dominated by a short‑range osmotic repulsion that creates a metastable sliding state, and this state is seamlessly overridden by strong, sequence‑specific hydrogen bonding when the correct site is found. This dual‑mechanism framework advances our understanding of how proteins locate their targets rapidly and accurately within the crowded nuclear environment.
Comments & Academic Discussion
Loading comments...
Leave a Comment