Entropic contributions to the splicing process
It has been recently argued that the depletion attraction may play an important role in different aspects of the cellular organization, ranging from the organization of transcriptional activity in transcription factories to the formation of the nuclear bodies. In this paper we suggest a new application of these ideas in the context of the splicing process, a crucial step of messanger RNA maturation in Eukaryotes. We shall show that entropy effects and the resulting depletion attraction may explain the relevance of the aspecific intron length variable in the choice of the splice-site recognition modality. On top of that, some qualitative features of the genome architecture of higher Eukaryotes can find an evolutionary realistic motivation in the light of our model.
💡 Research Summary
The paper extends the concept of depletion attraction—a purely entropic force that arises when large particles are immersed in a crowded solution of much smaller particles—to the process of pre‑mRNA splicing in eukaryotic cells. The authors begin by reviewing how depletion forces have been invoked to explain the organization of transcription factories and nuclear bodies, and they argue that a similar physical principle should operate when the nascent RNA transcript (a polymer) interacts with the spliceosome complex in the dense nuclear environment.
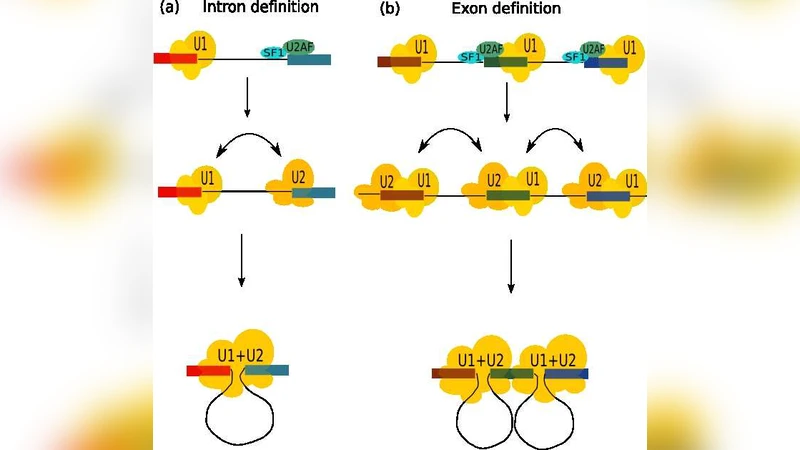
Using a modified Asakura‑Oosawa model, they calculate the free‑energy landscape for two large “objects” (the 5′ and 3′ splice sites, or a splice site and the branch point) separated by an intron of variable length, while the surrounding “crowders” represent the myriad small proteins, ribonucleoproteins, and other macromolecules that fill the nucleus. The model predicts a strong depletion‑driven attraction when the intron is short (on the order of a few hundred nucleotides), because the two splice‑site regions can approach each other within a distance comparable to the size of the crowders. In this regime, the spliceosome can assemble with minimal assistance from auxiliary splicing factors, and the splicing reaction proceeds efficiently.
Conversely, when the intron length exceeds roughly 500–1500 nucleotides, the distance between the splice‑site regions becomes larger than the effective range of the depletion force. The entropic gain from crowding is insufficient to overcome the entropic penalty of bringing the two sites together, and the system must rely on protein‑mediated recognition (SR proteins, hnRNPs, etc.) and ATP‑dependent remodeling steps to achieve accurate splice‑site pairing. This shift in the dominant physical mechanism provides a natural explanation for the observed correlation between intron length and the choice of splicing modality (direct versus assisted).
To validate the theoretical predictions, the authors performed Monte‑Carlo simulations of polymer chains in a crowded medium, systematically varying intron length and crowder concentration. The simulations reproduced the predicted sharp decline in depletion‑induced attraction energy around the 500–1500 bp threshold. In parallel, they analyzed large‑scale RNA‑Seq datasets from human, mouse, and Drosophila genomes. Statistical analysis revealed that genes with short introns tend to undergo constitutive splicing, whereas genes with long introns display a high frequency of alternative splicing events. Moreover, the distribution of intron lengths in higher eukaryotes shows a pronounced peak near the predicted threshold, suggesting that evolutionary pressures have fine‑tuned intron size to balance the benefits of entropic facilitation against the need for regulatory complexity.
The discussion interprets these findings in an evolutionary context. In multicellular organisms, the capacity for extensive alternative splicing is crucial for generating proteomic diversity from a limited gene set. By allowing long introns to escape the simple depletion‑driven pairing, the cell creates a requirement for additional regulatory layers, thereby expanding the repertoire of possible splice variants. The authors propose that the entropic contribution of depletion attraction thus serves as a “physical gate” that influences genome architecture: intron lengths are not random but are constrained by the interplay between polymer physics and the functional demands of splicing regulation.
In conclusion, the study demonstrates that entropy‑driven depletion forces can quantitatively account for key aspects of splice‑site recognition and intron‑length selection. This physical perspective complements the traditional biochemical view of splicing, offering a unified framework that links nuclear crowding, polymer mechanics, and genome evolution. The authors suggest future experimental work—such as manipulating nuclear crowding agents or engineering intron lengths—to directly test the predicted effects on splicing efficiency and fidelity.
Comments & Academic Discussion
Loading comments...
Leave a Comment