Intrinsic fluctuations in stochastic delay systems: theoretical description and application to a simple model of gene regulation
The effects of intrinsic noise on stochastic delay systems is studied within an expansion in the inverse system size. We show that the stochastic nature of the underlying dynamics may induce oscillatory behaviour in parameter ranges where the deterministic system does not sustain cycles, and compute the power spectra of these stochastic oscillations analytically, in good agreement with simulations. The theory is developed in the context of a simple one-dimensional toy model, but is applicable more generally. Gene regulatory systems in particular often contain only a small number of molecules, leading to significant fluctuations in mRNA and protein concentrations. As an application we therefore study a minimalistic model of the expression levels of hes1 mRNA and Hes1 protein, representing the simple motif of an auto-inhibitory feedback loop and motivated by its relevance to somite segmentation.
💡 Research Summary
The paper investigates how intrinsic stochasticity influences the dynamics of systems that contain explicit time delays. Using the van Kampen system‑size expansion, the authors separate the deterministic limit (obtained as the system size N → ∞) from the leading‑order stochastic corrections that scale as 1/N. In the deterministic limit the model reduces to a set of delay differential equations (DDEs) whose fixed points can be stable or, for certain parameter choices, support limit‑cycle oscillations. The novelty of the work lies in retaining the 1/N fluctuation term, which introduces a Poisson‑type noise that is itself delayed. Linearising around a deterministic fixed point yields a delayed linear stochastic differential equation whose characteristic equation possesses complex eigenvalues even when the deterministic Jacobian predicts purely real, negative eigenvalues. Consequently, the stochastic system exhibits “noise‑induced oscillations”: coherent, quasi‑periodic fluctuations that are absent from the deterministic description.
The authors derive an analytical expression for the power spectrum S(ω) of these oscillations. By Fourier‑transforming the linearised stochastic equations, they obtain a rational function whose denominator contains both ω‑dependent terms from the usual diffusion matrix and trigonometric contributions arising from the delay τ. The spectrum displays a pronounced peak at a frequency ω* that depends on the delay and on the kinetic rates of production and degradation. The peak’s width scales inversely with N, confirming that larger systems exhibit weaker, broader fluctuations, while small systems (low molecule numbers) show sharp, high‑amplitude peaks. The analytical predictions are validated against Gillespie‑type stochastic simulations that incorporate explicit delays; the agreement is excellent across a wide range of parameters.
To illustrate the generality of the approach, the authors first apply it to a minimal one‑dimensional “toy” model consisting of a single species X that is produced after a fixed delay τ and degraded instantaneously. In the deterministic limit the system settles to a stable fixed point, but the stochastic analysis predicts a clear spectral peak, which is reproduced by delayed Gillespie simulations. This example demonstrates that even the simplest delayed birth‑death process can generate coherent stochastic oscillations solely due to intrinsic noise.
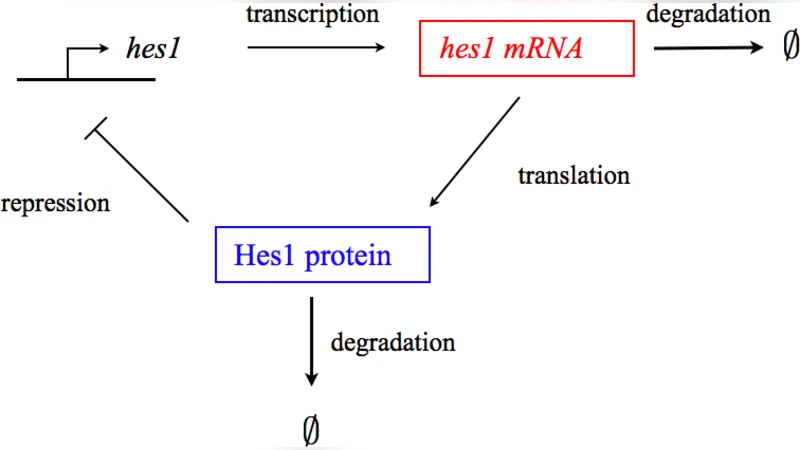
The second, biologically motivated application focuses on the Hes1 gene regulatory circuit, a classic example of an auto‑inhibitory feedback loop that controls somite segmentation during vertebrate development. The model includes four reactions: transcription of Hes1 mRNA, translation into Hes1 protein, degradation of mRNA, and degradation of protein. Crucially, the transcriptional repression is delayed: the protein produced at time t suppresses mRNA synthesis at time t + τ, reflecting the time required for transcription, splicing, and nuclear transport. Parameter values (transcription rate, translation rate, degradation constants, and τ≈30 min) are taken from experimental literature, and the typical copy numbers are low (10–100 molecules), placing the system squarely in the regime where intrinsic noise is significant.
Deterministic analysis of this delayed feedback system predicts a stable steady state with no oscillations. However, when the 1/N stochastic correction is included, the linearised system acquires a pair of complex conjugate eigenvalues, leading to a power‑spectral peak around a frequency corresponding to a period of roughly 2 hours—precisely the oscillation period observed experimentally for Hes1 protein and mRNA levels in cultured cells. Simulations using the delayed Gillespie algorithm reproduce this oscillatory behaviour, and the analytically derived spectrum matches the simulated one quantitatively. This result provides a compelling mechanistic explanation: the combination of a delayed negative feedback loop and intrinsic molecular noise is sufficient to generate the experimentally observed Hes1 oscillations, without invoking additional regulatory components or external forcing.
The paper concludes by emphasizing several broader implications. First, it shows that delay and noise are not merely additive perturbations; their interaction can fundamentally alter system dynamics, turning a deterministically stable system into a stochastic oscillator. Second, the analytical power‑spectral formula offers a practical tool for parameter inference: by fitting experimental spectra one can estimate otherwise inaccessible quantities such as effective delay times or kinetic rates. Third, although the study focuses on a single‑species toy model and a two‑species gene circuit, the methodology extends naturally to higher‑dimensional networks with multiple delays, making it relevant for a wide class of synthetic and natural biochemical circuits. Finally, the work highlights the importance of stochastic modeling in contexts where molecule numbers are low, as is typical for gene regulation, and suggests that many biological rhythms may arise from the interplay of intrinsic noise and delayed feedback rather than from deterministic limit‑cycle mechanisms alone.
Comments & Academic Discussion
Loading comments...
Leave a Comment